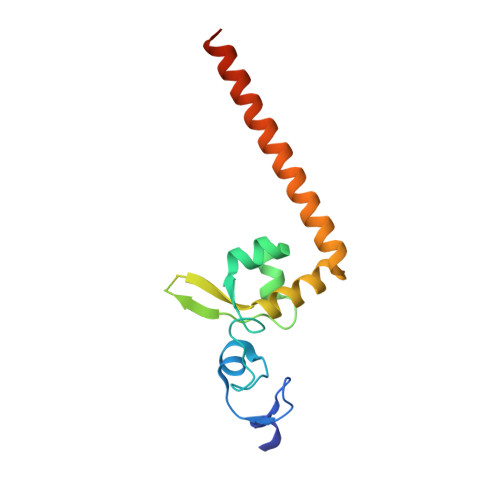
Structural characterization of the redefined DNA-binding domain of human XPA.
Lian, F.M., Yang, X., Yang, W., Jiang, Y.L., Qian, C.(2019) Biochem Biophys Res Commun 514: 985-990
- PubMed: 31092331
- DOI: https://doi.org/10.1016/j.bbrc.2019.05.050
- Primary Citation Related Structures:
6J44 - PubMed Abstract:
XPA (xeroderma pigmentosum complementation group A), a key scaffold protein in nucleotide excision repair (NER) pathway, is important in DNA damage verification and repair proteins recruitment. Earlier studies had mapped the minimal DNA-binding domain (MBD) of XPA to a region corresponding to residues 98-219. However, recent studies indicated that the region involving residues 98-239 is the redefined DNA-binding domain (DBD), which binds to DNA substrates with a much higher binding affinity than MBD and possesses a nearly identical binding affinity to the full-length XPA protein. However, the structure of the redefined DBD domain of XPA (XPA-DBD) remains to be investigated. Here, we present the crystal structure of XPA-DBD at 2.06 Å resolution. Structure of the C-terminal region of XPA has been extended by 21 residues (Arg211-Arg231) as compared with previously reported MBD structures. The structure reveals that the C-terminal extension (Arg211-Arg231) is folded as an α-helix with multiple basic residues. The positively charged surface formed in the last C-terminal helix suggests its critical role in DNA binding. Further structural analysis demonstrates that the last C-terminal region (Asp217-Thr239) of XPA-DBD might undergo a conformational change to directly bind to the DNA substrates. This study provides a structural basis for understanding the possible mechanism of enhanced DNA-binding affinity of XPA-DBD.
- Key Laboratory of Precision Oncology of Shandong Higher Education, Institute of Precision Medicine, Jining Medical University, Jining, Shandong, 272067, China. Electronic address: fmlian@mail.jnmc.edu.cn.
Organizational Affiliation: