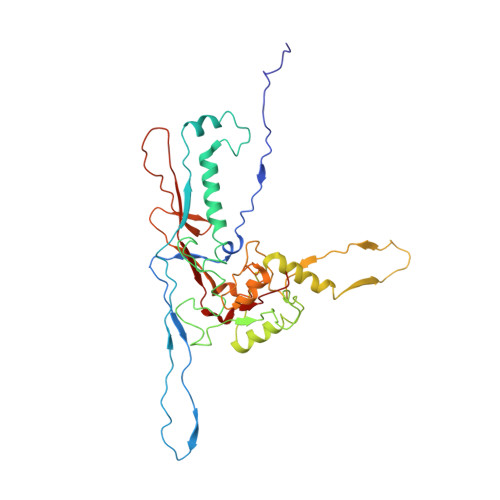
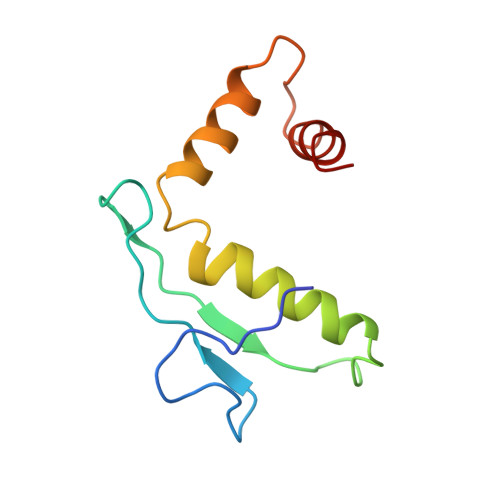
Capsid Structure of a Freshwater Cyanophage Siphoviridae Mic1.
Jin, H., Jiang, Y.L., Yang, F., Zhang, J.T., Li, W.F., Zhou, K., Ju, J., Chen, Y., Zhou, C.Z.(2019) Structure 27: 1508
- PubMed: 31378451
- DOI: https://doi.org/10.1016/j.str.2019.07.003
- Primary Citation Related Structures:
6J3Q - PubMed Abstract:
Cyanobacteria are the most abundant photosynthetic microorganisms, the global distribution of which is mainly regulated by the corresponding cyanophages. A systematic screening of water samples in the Lake Chaohu enabled us to isolate a freshwater siphocyanophage that infects Microcystis wesenbergii, thus termed Mic1. Using cryoelectron microscopy, we solved the 3.5-Å structure of Mic1 capsid. The major capsid protein gp40 of an HK97-like fold forms two types of capsomers, hexons and pentons. The capsomers interact with each other via the interweaved N-terminal arms of gp40 in addition to a tail-in-mouth joint along the three-fold symmetric axis, resulting in the assembly of capsid in a mortise-and-tenon pattern. The novel-fold cement protein gp47 sticks at the two-fold symmetric axis and further fixes the capsid. These findings provide structural insights into the assembly of cyanophages, and set up a platform to explore the mechanism of specific interactions and co-evolution with cyanobacteria.
- Hefei National Laboratory for Physical Sciences at the Microscale and School of Life Sciences, University of Science and Technology of China, Hefei, Anhui 230026, China.
Organizational Affiliation: