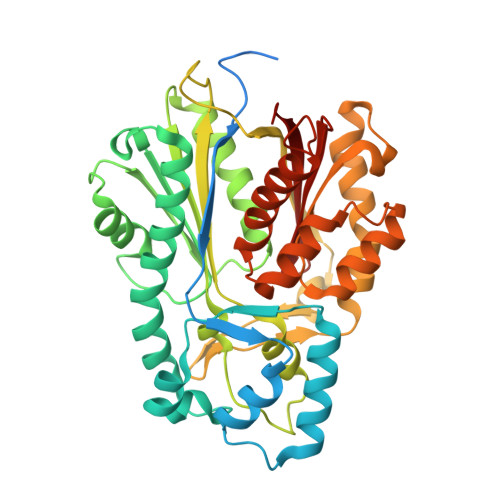
Tropane alkaloids biosynthesis involves an unusual type III polyketide synthase and non-enzymatic condensation.
Huang, J.P., Fang, C., Ma, X., Wang, L., Yang, J., Luo, J., Yan, Y., Zhang, Y., Huang, S.X.(2019) Nat Commun 10: 4036-4036
- PubMed: 31492848
- DOI: https://doi.org/10.1038/s41467-019-11987-z
- Primary Citation of Related Structures:
6J1M, 6J1N - PubMed Abstract:
The skeleton of tropane alkaloids is derived from ornithine-derived N-methylpyrrolinium and two malonyl-CoA units. The enzymatic mechanism that connects N-methylpyrrolinium and malonyl-CoA units remains unknown. Here, we report the characterization of three pyrrolidine ketide synthases (PYKS), AaPYKS, DsPYKS, and AbPYKS, from three different hyoscyamine- and scopolamine-producing plants. By examining the crystal structure and biochemical activity of AaPYKS, we show that the reaction mechanism involves PYKS-mediated malonyl-CoA condensation to generate a 3-oxo-glutaric acid intermediate that can undergo non-enzymatic Mannich-like condensation with N-methylpyrrolinium to yield the racemic 4-(1-methyl-2-pyrrolidinyl)-3-oxobutanoic acid. This study therefore provides a long sought-after biosynthetic mechanism to explain condensation between N-methylpyrrolinium and acetate units and, more importantly, identifies an unusual plant type III polyketide synthase that can only catalyze one round of malonyl-CoA condensation.
- State Key Laboratory of Phytochemistry and Plant Resources in West China, and CAS Center for Excellence in Molecular Plant Sciences, Kunming Institute of Botany, Chinese Academy of Sciences, Kunming, 650201, China.
Organizational Affiliation: