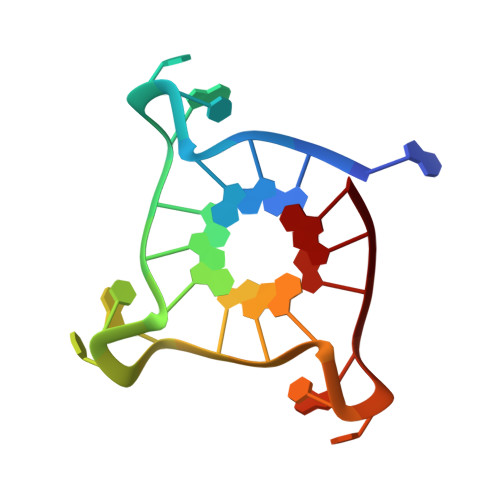
Probing G-quadruplex topologies and recognition concurrently in real time and 3D using a dual-app nucleoside probe.
Nuthanakanti, A., Ahmed, I., Khatik, S.Y., Saikrishnan, K., Srivatsan, S.G.(2019) Nucleic Acids Res 47: 6059-6072
- PubMed: 31106340 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkz419
- Primary Citation Related Structures:
6IP3, 6IP7, 6ISW - PubMed Abstract:
Comprehensive understanding of structure and recognition properties of regulatory nucleic acid elements in real time and atomic level is highly important to devise efficient therapeutic strategies. Here, we report the establishment of an innovative biophysical platform using a dual-app nucleoside analog, which serves as a common probe to detect and correlate different GQ structures and ligand binding under equilibrium conditions and in 3D by fluorescence and X-ray crystallography techniques. The probe (SedU) is composed of a microenvironment-sensitive fluorophore and an excellent anomalous X-ray scatterer (Se), which is assembled by attaching a selenophene ring at 5-position of 2'-deoxyuridine. SedU incorporated into the loop region of human telomeric DNA repeat fluorescently distinguished subtle differences in GQ topologies and enabled quantify ligand binding to different topologies. Importantly, anomalous X-ray dispersion signal from Se could be used to determine the structure of GQs. As the probe is minimally perturbing, a direct comparison of fluorescence data and crystal structures provided structural insights on how the probe senses different GQ conformations without affecting the native fold. Taken together, our dual-app probe represents a new class of tool that opens up new experimental strategies to concurrently investigate nucleic acid structure and recognition in real time and 3D.
- Department of Chemistry, Indian Institute of Science Education and Research (IISER), Pune, Dr. Homi Bhabha Road, Pune 411008, India.
Organizational Affiliation: