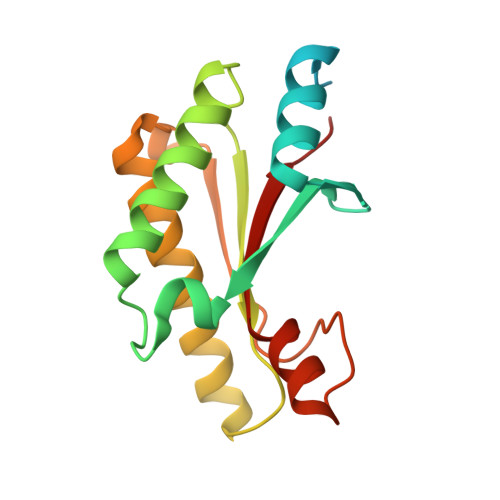
Structural analysis of activating mutants of YfiB from Pseudomonas aeruginosa PAO1.
Li, S., Li, T., Teng, X., Lou, X., Xu, Y., Zhang, Q., Bartlam, M.(2018) Biochem Biophys Res Commun 506: 997-1003
- PubMed: 30404734
- DOI: https://doi.org/10.1016/j.bbrc.2018.10.190
- Primary Citation Related Structures:
6IKI, 6IKJ, 6IKK - PubMed Abstract:
Bacterial cyclic-di-GMP (c-di-GMP) is an important messenger molecule that influences diverse cellular processes including motility, virulence and cytotoxicity systems, polysaccharide synthesis and biofilm formation. The YfiBNR tripartite signalling system in P. aeruginosa modulates the cellular c-di-GMP levels in response to signals received from the periplasm. In this study, we analyse the structures of activating mutants of the outer membrane protein YfiB that give rise to increased surface attachment and biofilm formation. The F48S and W55L mutants of YfiB(27-168) crystallize in the same dimeric arrangement as our previously reported YfiB structures that preclude complex formation with YfiR. The L43P mutant of YfiB(27-168) is monomeric and forms a stable complex with YfiR. The YfiB(L43P)-YfiR crystal structure reveals a dramatic rearrangement of the N-terminal fragment, which is implicated in increased YfiB activation and membrane attachment, upon YfiR binding. Comparison with our previous complex structure between YfiB(59-168) and YfiR reveals extensive interactions between the N-terminal fragment of YfiB (residues 35-55) and YfiR.
- State Key Laboratory of Medicinal Chemical Biology, Nankai University, Tianjin, 300071, People's Republic of China; College of Life Sciences, Nankai University, Tianjin, 300071, People's Republic of China.
Organizational Affiliation: