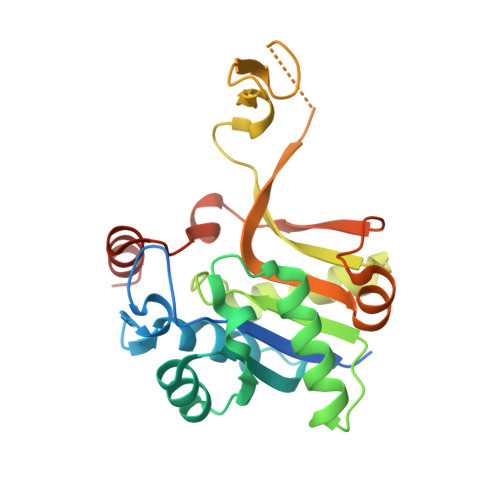
Structural and functional characterization of CMP-N-acetylneuraminate synthetase from Vibrio cholerae.
Bose, S., Purkait, D., Joseph, D., Nayak, V., Subramanian, R.(2019) Acta Crystallogr D Struct Biol 75: 564-577
- PubMed: 31205019 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2059798319006831
- Primary Citation Related Structures:
6IFD, 6IFI - PubMed Abstract:
Several pathogenic bacteria utilize sialic acid, including host-derived N-acetylneuraminic acid (Neu5Ac), in at least two ways: they use it as a nutrient source and as a host-evasion strategy by coating themselves with Neu5Ac. Given the significant role of sialic acid in pathogenesis and host-gut colonization by various pathogenic bacteria, including Neisseria meningitidis, Haemophilus influenzae, Pasteurella multocida and Vibrio cholerae, several enzymes of the sialic acid catabolic, biosynthetic and incorporation pathways are considered to be potential drug targets. In this work, findings on the structural and functional characterization of CMP-N-acetylneuraminate synthetase (CMAS), a key enzyme in the incorporation pathway, from Vibrio cholerae are reported. CMAS catalyzes the synthesis of CMP-sialic acid by utilizing CTP and sialic acid. Crystal structures of the apo and the CDP-bound forms of the enzyme were determined, which allowed the identification of the metal cofactor Mg 2+ in the active site interacting with CDP and the invariant Asp215 residue. While open and closed structural forms of the enzyme from eukaryotic and other bacterial species have already been characterized, a partially closed structure of V. cholerae CMAS (VcCMAS) observed upon CDP binding, representing an intermediate state, is reported here. The kinetic data suggest that VcCMAS is capable of activating the two most common sialic acid derivatives, Neu5Ac and Neu5Gc. Amino-acid sequence and structural comparison of the active site of VcCMAS with those of eukaryotic and other bacterial counterparts reveal a diverse hydrophobic pocket that interacts with the C5 substituents of sialic acid. Analyses of the thermodynamic signatures obtained from the binding of the nucleotide (CTP) and the product (CMP-sialic acid) to VcCMAS provide fundamental information on the energetics of the binding process.
- Institute for Stem Cell Science and Regenerative Medicine, GKVK Post, Bellary Road, Bangalore 560 065, India.
Organizational Affiliation: