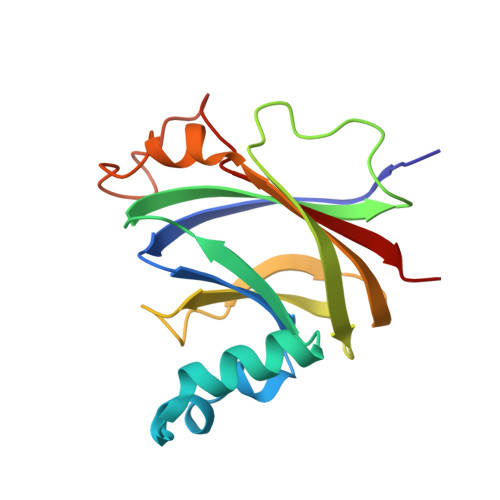
The structure of CgnJ, a domain of unknown function protein from the crocagin gene cluster.
Adam, S., Klein, A., Surup, F., Koehnke, J.(2019) Acta Crystallogr F Struct Biol Commun 75: 205-211
- PubMed: 30839296
- DOI: https://doi.org/10.1107/S2053230X19000712
- Primary Citation of Related Structures:
6I86 - PubMed Abstract:
Natural products often contain interesting new chemical entities that are introduced into the structure of a compound by the enzymatic machinery of the producing organism. The recently described crocagins are novel polycyclic peptides which belong to the class of ribosomally synthesized and post-translationally modified peptide natural products. They have been shown to bind to the conserved prokaryotic carbon-storage regulator A in vitro. In efforts to understand crocagin biosynthesis, the putative biosynthetic genes were expressed and purified. Here, the first crystal structure of a protein from the crocagin-biosynthetic gene cluster, CgnJ, a domain of unknown function protein, is reported. Possible functions of this protein were explored by structural and sequence homology analyses. Even though the sequence homology to proteins in the Protein Data Bank is low, the protein shows significant structural homology to a protein with known function within the competency system of Bacillus subtilis, ComJ, leading to the hypothesis of a similar role of the protein within the producing organism.
- Structural Biology of Biosynthetic Enzymes, Helmholtz Institute for Pharmaceutical Research Saarland, Universität des Saarlandes Gebäude E8.1, 66123 Saarbrücken, Germany.
Organizational Affiliation: