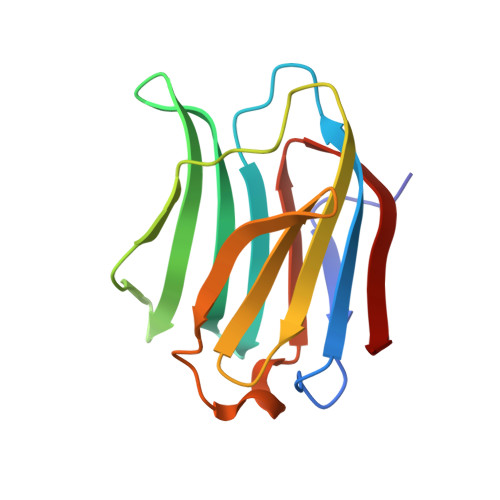
Substituted polyfluoroaryl interactions with an arginine side chain in galectin-3 are governed by steric-, desolvation and electronic conjugation effects.
Kumar, R., Peterson, K., Misini Ignjatovic, M., Leffler, H., Ryde, U., Nilsson, U.J., Logan, D.T.(2019) Org Biomol Chem 17: 1081-1089
- PubMed: 30632578 Search on PubMed
- DOI: https://doi.org/10.1039/c8ob02888e
- Primary Citation Related Structures:
6I74, 6I75, 6I76, 6I77, 6I78 - PubMed Abstract:
In the β-d-galactopyranoside-binding protein galectin-3, synthetic inhibitors substituted at the 3-position of a thiodigalactoside core cause the formation of an aglycone binding pocket through the displacement of an arginine residue (Arg144) from its position in the apoprotein. To examine in detail the role of different molecular interactions in this pocket, we have synthesized a series of nine 3-(4-(2,3,5,6-tetrafluorophenyl)-1,2,3-triazol-1-yl)-thiogalactosides with different para substituents and measured their affinities to galectin-3 using a fluorescence polarization assay. High-resolution crystal structures (<1.3 Å) have been determined for five of the ligands in complex with the C-terminal domain of galectin-3. The binding affinities are rationalised with the help of the three-dimensional structures and quantum-mechanical calculations. Three effects seem to be involved: Firstly, the binding pocket is too small for the largest ligands with ethyl and methyl. Secondly, for the other ligands, the affinity seems to be determined mainly by desolvation effects, disfavouring the polar substituents, but this is partly counteracted by the cation-π interaction with Arg144, which stacks on top of the substituted tetrafluorophenyl group in all complexes. The results provide detailed insight into interactions of fluorinated phenyl moieties with arginine-containing protein binding sites and the complex interplay of different energetic components in defining the binding affinity.
- Biochemistry and Structural Biology, Centre for Molecular Protein Science, Department of Chemistry, Lund University, Box 124, SE-221 00 Lund, Sweden. derek.logan@biochemistry.lu.se.
Organizational Affiliation: