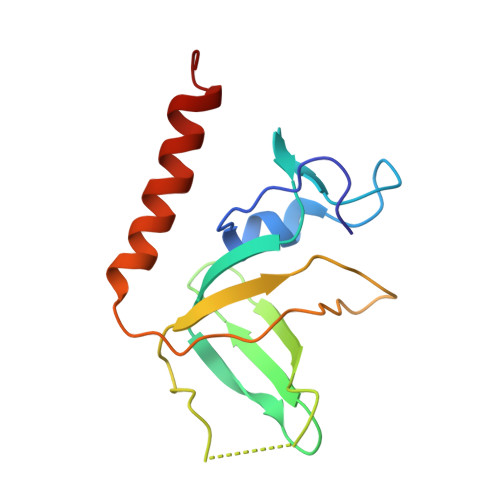
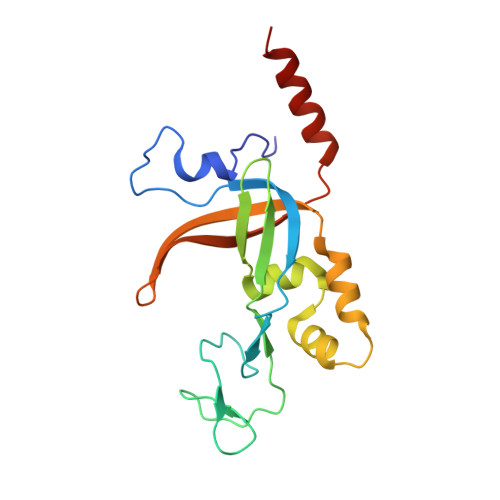
A structural and dynamic model for the assembly of Replication Protein A on single-stranded DNA.
Yates, L.A., Aramayo, R.J., Pokhrel, N., Caldwell, C.C., Kaplan, J.A., Perera, R.L., Spies, M., Antony, E., Zhang, X.(2018) Nat Commun 9: 5447-5447
- PubMed: 30575763 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-018-07883-7
- Primary Citation Related Structures:
6I52 - PubMed Abstract:
Replication Protein A (RPA), the major eukaryotic single stranded DNA-binding protein, binds to exposed ssDNA to protect it from nucleases, participates in a myriad of nucleic acid transactions and coordinates the recruitment of other important players. RPA is a heterotrimer and coats long stretches of single-stranded DNA (ssDNA). The precise molecular architecture of the RPA subunits and its DNA binding domains (DBDs) during assembly is poorly understood. Using cryo electron microscopy we obtained a 3D reconstruction of the RPA trimerisation core bound with ssDNA (∼55 kDa) at ∼4.7 Å resolution and a dimeric RPA assembly on ssDNA. FRET-based solution studies reveal dynamic rearrangements of DBDs during coordinated RPA binding and this activity is regulated by phosphorylation at S178 in RPA70. We present a structural model on how dynamic DBDs promote the cooperative assembly of multiple RPAs on long ssDNA.
- Section of Structural Biology, Department of Medicine, Imperial College London, Sir Alexander Fleming Building, South Kensington, London, SW7 2AZ, UK.
Organizational Affiliation: