Discovery and Characterization of the Potent and Selective P2X4 InhibitorN-[4-(3-Chlorophenoxy)-3-sulfamoylphenyl]-2-phenylacetamide (BAY-1797) and Structure-Guided Amelioration of Its CYP3A4 Induction Profile.
Werner, S., Mesch, S., Hillig, R.C., Ter Laak, A., Klint, J., Neagoe, I., Laux-Biehlmann, A., Dahllof, H., Brauer, N., Puetter, V., Nubbemeyer, R., Schulz, S., Bairlein, M., Zollner, T.M., Steinmeyer, A.(2019) J Med Chem 62: 11194-11217
- PubMed: 31746599 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.9b01304
- Primary Citation Related Structures:
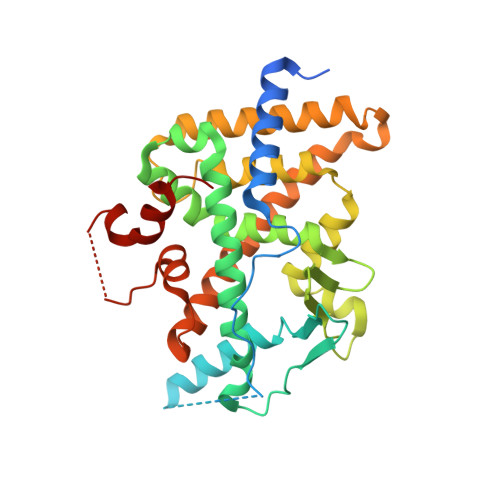
6HTY - PubMed Abstract:
The P2X4 receptor is a ligand-gated ion channel that is expressed on a variety of cell types, especially those involved in inflammatory and immune processes. High-throughput screening led to a new class of P2X4 inhibitors with substantial CYP 3A4 induction in human hepatocytes. A structure-guided optimization with respect to decreased pregnane X receptor (PXR) binding was started. It was found that the introduction of larger and more polar substituents on the ether linker led to less PXR binding while maintaining the P2X4 inhibitory potency. This translated into significantly reduced CYP 3A4 induction for compounds 71 and 73 . Unfortunately, the in vivo pharmacokinetic (PK) profiles of these compounds were insufficient for the desired profile in humans. However, BAY-1797 ( 10 ) was identified and characterized as a potent and selective P2X4 antagonist. This compound is suitable for in vivo studies in rodents, and the anti-inflammatory and anti-nociceptive effects of BAY-1797 were demonstrated in a mouse complete Freund's adjuvant (CFA) inflammatory pain model.
- Bayer AG, Research & Development, Pharmaceuticals , 13353 Berlin , Germany.
Organizational Affiliation: