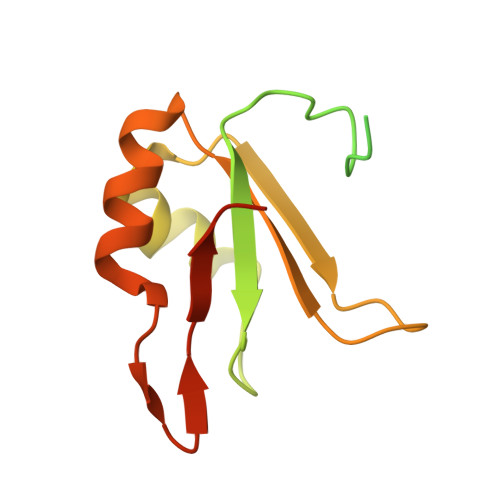
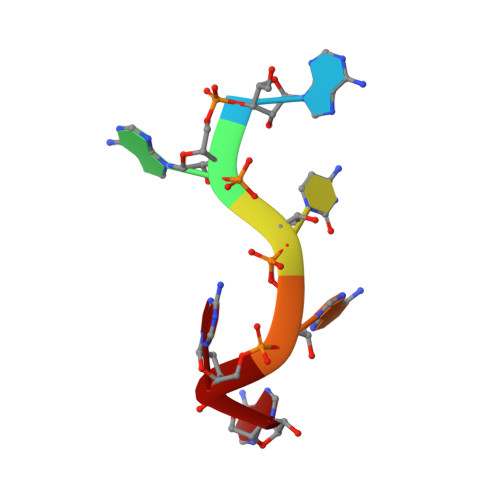
Structure of SRSF1 RRM1 bound to RNA reveals an unexpected bimodal mode of interaction and explains its involvement in SMN1 exon7 splicing.
Clery, A., Krepl, M., Nguyen, C.K.X., Moursy, A., Jorjani, H., Katsantoni, M., Okoniewski, M., Mittal, N., Zavolan, M., Sponer, J., Allain, F.H.(2021) Nat Commun 12: 428-428
- PubMed: 33462199 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-20481-w
- Primary Citation Related Structures:
6HPJ - PubMed Abstract:
The human prototypical SR protein SRSF1 is an oncoprotein that contains two RRMs and plays a pivotal role in RNA metabolism. We determined the structure of the RRM1 bound to RNA and found that the domain binds preferentially to a CN motif (N is for any nucleotide). Based on this solution structure, we engineered a protein containing a single glutamate to asparagine mutation (E87N), which gains the ability to bind to uridines and thereby activates SMN exon7 inclusion, a strategy that is used to cure spinal muscular atrophy. Finally, we revealed that the flexible inter-RRM linker of SRSF1 allows RRM1 to bind RNA on both sides of RRM2 binding site. Besides revealing an unexpected bimodal mode of interaction of SRSF1 with RNA, which will be of interest to design new therapeutic strategies, this study brings a new perspective on the mode of action of SRSF1 in cells.
- Department of Biology, Institute of Biochemistry, ETH Zurich, Zurich, Switzerland. aclery@mol.biol.ethz.ch.
Organizational Affiliation: