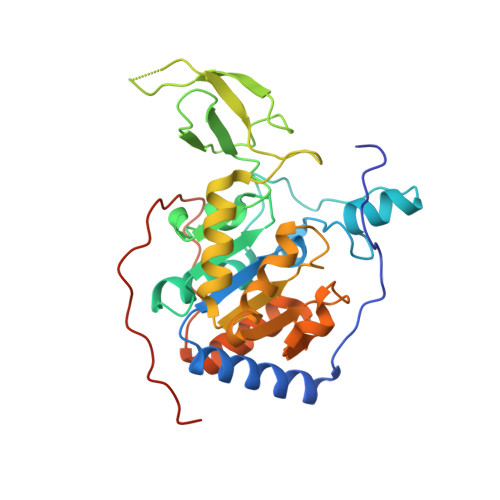
Structural Basis of Sirtuin 6 Inhibition by the Hydroxamate Trichostatin A: Implications for Protein Deacylase Drug Development.
You, W., Steegborn, C.(2018) J Med Chem 61: 10922-10928
- PubMed: 30395713 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.8b01455
- Primary Citation Related Structures:
6HOY - PubMed Abstract:
Protein lysine deacylases comprise three zinc-dependent families and the NAD + -dependent sirtuins Sirt1-7, which contribute to aging-related diseases. Few Sirt6-specific inhibitors are available. Trichostatin A, which belongs to the potent, zinc-chelating hydroxamate inhibitors of zinc-dependent deacylases, was recently found to potently and isoform-specifically inhibit Sirt6. We solved a crystal structure of a Sirt6/ADP-ribose/trichostatin A complex, which reveals nicotinamide pocket and acyl channel as binding site and provides interaction details supporting the development of improved deacylase inhibitors.
- Department of Biochemistry , University of Bayreuth , Universitätsstraße 30 , 95445 Bayreuth , Germany.
Organizational Affiliation: