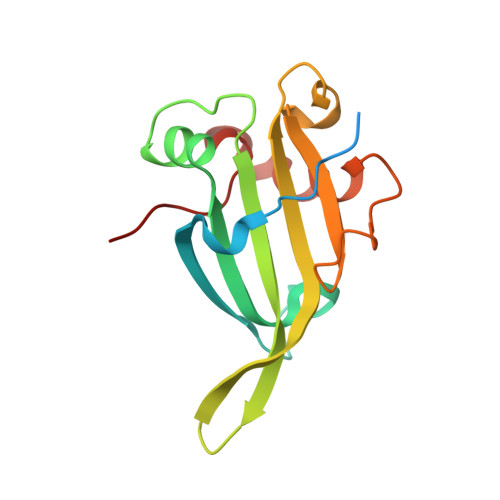
One fold, two functions: cytochrome P460 and cytochromec'-beta from the methanotrophMethylococcus capsulatus(Bath).
Adams, H.R., Krewson, C., Vardanega, J.E., Fujii, S., Moreno, T., Sambongi, Y., Svistunenko, D., Paps, J., Andrew, C.R., Hough, M.A.(2019) Chem Sci 10: 3031-3041
- PubMed: 30996884 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1039/c8sc05210g
- Primary Citation Related Structures:
6HIH, 6HIU - PubMed Abstract:
Nature is adept at utilising highly similar protein folds to carry out very different functions, yet the mechanisms by which this functional divergence occurs remain poorly characterised. In certain methanotrophic bacteria, two homologous pentacoordinate c-type heme proteins have been identified: a cytochrome P460 (cyt P460) and a cytochrome c '-β (cyt cp-β). Cytochromes P460 are able to convert hydroxylamine to nitrous oxide (N 2 O), a potent greenhouse gas. This reactivity is similar to that of hydroxylamine oxidoreductase (HAO), which is a key enzyme in nitrifying and methanotrophic bacteria. Cyt P460 and HAO both have unusual protein-heme cross-links, formed by a Tyr residue in HAO and a Lys in cyt P460. In contrast, cyts cp-β (the only known cytochromes c ' with a β-sheet fold) lack this crosslink and appears to be optimized for binding non-polar molecules (including NO and CO) without enzymatic conversion. Our bioinformatics analysis supports the proposal that cyt cp-β may have evolved from cyt P460 via a gene duplication event. Using high-resolution X-ray crystallography, UV-visible absorption, electron paramagnetic resonance (EPR) and resonance Raman spectroscopy, we have characterized the overall protein folding and active site structures of cyt cp-β and cyt P460 from the obligate methanotroph, Methylococcus capsulatus (Bath). These proteins display a similar β-sheet protein fold, together with a pattern of changes to the heme pocket regions and localised tertiary structure that have converted a hydroxylamine oxidizing enzyme into a gas-binding protein. Structural comparisons provide insights relevant to enzyme redesign for synthetic enzymology and engineering of gas sensor proteins. We also show the widespread occurrence of cyts cp-β and characterise their phylogeny.
- School of Biological Sciences , University of Essex , Wivenhoe Park , Colchester , Essex CO4 3SQ , UK . Email: mahough@essex.ac.uk.
Organizational Affiliation: