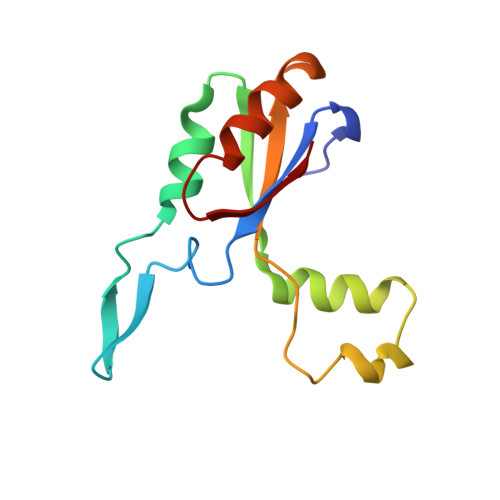
Structure of the Large Extracellular Loop of FtsX and Its Interaction with the Essential Peptidoglycan Hydrolase PcsB in Streptococcus pneumoniae.
Rued, B.E., Alcorlo, M., Edmonds, K.A., Martinez-Caballero, S., Straume, D., Fu, Y., Bruce, K.E., Wu, H., Havarstein, L.S., Hermoso, J.A., Winkler, M.E., Giedroc, D.P.(2019) mBio 10
- PubMed: 30696736 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/mBio.02622-18
- Primary Citation Related Structures:
6HE6, 6HEE, 6HFX, 6MK7 - PubMed Abstract:
Streptococcus pneumoniae is a leading killer of infants and immunocompromised adults and has become increasingly resistant to major antibiotics. Therefore, the development of new antibiotic strategies is desperately needed. Targeting bacterial cell division is one such strategy, specifically by targeting proteins that are essential for the synthesis and breakdown of peptidoglycan. One complex important to this process is FtsEX. FtsEX comprises a cell division-regulating integral membrane protein (FtsX) and a cytoplasmic ATPase (FtsE) that resembles an ATP-binding cassette (ABC) transporter. Here, we present nuclear magnetic resonance (NMR) solution structural and crystallographic models of the large extracellular domain of FtsX, denoted extracellular loop 1 (ECL1). The structure of ECL1 reveals an upper extended β-hairpin and a lower α-helical lobe, each extending from a mixed α-β core. The helical lobe mediates a physical interaction with the peptidoglycan hydrolase PcsB via the coiled-coil domain of PcsB (PscB CC ). Characterization of S. pneumoniae strain D39-derived strains harboring mutations in the α-helical lobe shows that this subdomain is essential for cell viability and required for proper cell division of S. pneumoniae IMPORTANCE FtsX is a ubiquitous bacterial integral membrane protein involved in cell division that regulates the activity of peptidoglycan (PG) hydrolases. FtsX is representative of a large group of ABC3 superfamily proteins that function as "mechanotransmitters," proteins that relay signals from the inside to the outside of the cell. Here, we present a structural characterization of the large extracellular loop, ECL1, of FtsX from the opportunistic human pathogen S. pneumoniae We show the molecular nature of the direct interaction between the peptidoglycan hydrolase PcsB and FtsX and demonstrate that this interaction is essential for cell viability. As such, FtsX represents an attractive, conserved target for the development of new classes of antibiotics.
- Department of Biology, Indiana University Bloomington, Bloomington, Indiana, USA.
Organizational Affiliation: