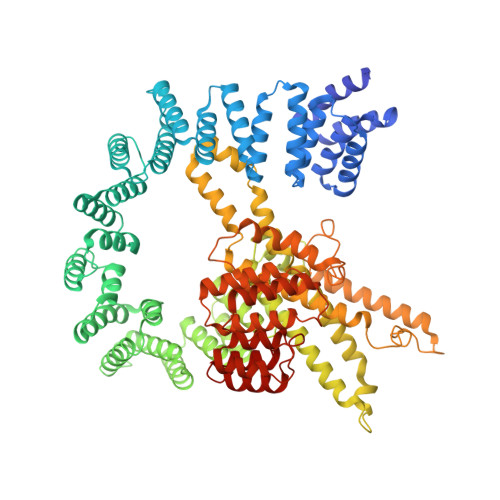
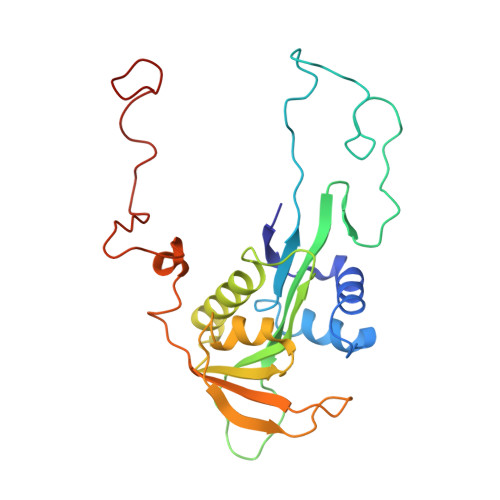
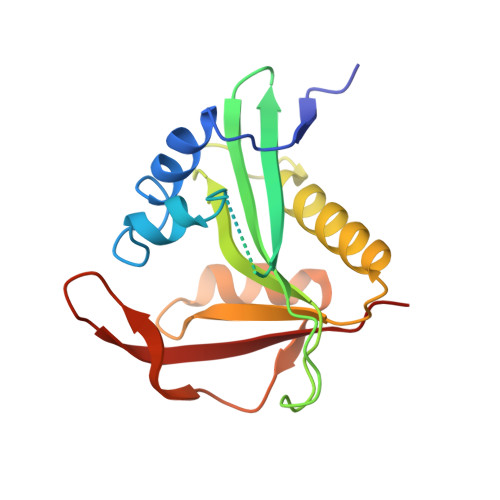
Ribosome-NatA architecture reveals that rRNA expansion segments coordinate N-terminal acetylation.
Knorr, A.G., Schmidt, C., Tesina, P., Berninghausen, O., Becker, T., Beatrix, B., Beckmann, R.(2019) Nat Struct Mol Biol 26: 35-39
- PubMed: 30559462 Search on PubMed
- DOI: https://doi.org/10.1038/s41594-018-0165-y
- Primary Citation Related Structures:
6HD5, 6HD7 - PubMed Abstract:
The majority of eukaryotic proteins are N-terminally α-acetylated by N-terminal acetyltransferases (NATs). Acetylation usually occurs co-translationally and defects have severe consequences. Nevertheless, it is unclear how these enzymes act in concert with the translating ribosome. Here, we report the structure of a native ribosome-NatA complex from Saccharomyces cerevisiae. NatA (comprising Naa10, Naa15 and Naa50) displays a unique mode of ribosome interaction by contacting eukaryotic-specific ribosomal RNA expansion segments in three out of four binding patches. Thereby, NatA is dynamically positioned directly underneath the ribosomal exit tunnel to facilitate modification of the emerging nascent peptide chain. Methionine amino peptidases, but not chaperones or signal recognition particle, would be able to bind concomitantly. This work assigns a function to the hitherto enigmatic ribosomal RNA expansion segments and provides mechanistic insights into co-translational protein maturation by N-terminal acetylation.
- Gene Center and Center for Integrated Protein Science Munich, Department of Biochemistry, University of Munich, Munich, Germany.
Organizational Affiliation: