Crystal structures of murine norovirus protruding domain with conjugated bile acid reveal a novel binding pocket.
Kilic, T., Koromyslova, A., Malak, V., Hansman, G.S.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Capsid protein | 303 | Murine norovirus GV/CR10/2005/USA | Mutation(s): 0 | 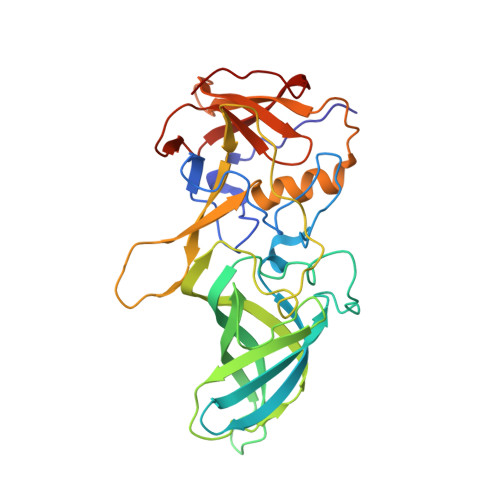 | |
UniProt | |||||
Find proteins for A7YK37 (Murine norovirus GV/CR10/2005/USA) Explore A7YK37 Go to UniProtKB: A7YK37 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | A7YK37 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| CMRF35-like molecule 1 | 111 | Mus musculus | Mutation(s): 0 Gene Names: Cd300lf, Clm1, Lmir3 |  | |
UniProt | |||||
Find proteins for Q6SJQ7 (Mus musculus) Explore Q6SJQ7 Go to UniProtKB: Q6SJQ7 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q6SJQ7 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 5 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| CHO Query on CHO | G [auth A], M [auth B], S [auth C], X [auth D] | GLYCOCHENODEOXYCHOLIC ACID C26 H43 N O5 GHCZAUBVMUEKKP-GYPHWSFCSA-N |  | ||
| PGE Query on PGE | EA [auth F] | TRIETHYLENE GLYCOL C6 H14 O4 ZIBGPFATKBEMQZ-UHFFFAOYSA-N |  | ||
| PEG Query on PEG | CA [auth E] | DI(HYDROXYETHYL)ETHER C4 H10 O3 MTHSVFCYNBDYFN-UHFFFAOYSA-N |  | ||
| EDO Query on EDO | AA [auth D] BA [auth E] DA [auth F] H [auth A] I [auth A] | 1,2-ETHANEDIOL C2 H6 O2 LYCAIKOWRPUZTN-UHFFFAOYSA-N |  | ||
| NA Query on NA | L [auth A], N [auth B], W [auth C], Y [auth D] | SODIUM ION Na FKNQFGJONOIPTF-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 75.94 | α = 90 |
| b = 137.55 | β = 95.84 |
| c = 93.8 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| XDS | data reduction |
| XSCALE | data scaling |
| PHASER | phasing |