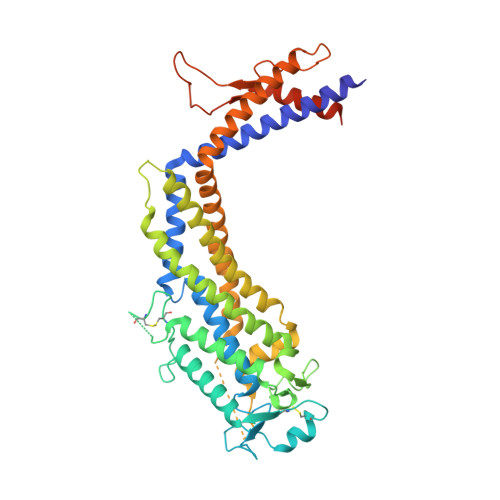
Hop-family Helicobacter outer membrane adhesins form a novel class of Type 5-like secretion proteins with an interrupted beta-barrel domain.
Coppens, F., Castaldo, G., Debraekeleer, A., Subedi, S., Moonens, K., Lo, A., Remaut, H.(2018) Mol Microbiol 110: 33-46
- PubMed: 29995350
- DOI: https://doi.org/10.1111/mmi.14075
- Primary Citation of Related Structures:
6GW5 - PubMed Abstract:
The human stomach pathogen Helicobacter pyloriattaches to healthy and inflamed gastric tissue through members of a paralogous family of 'Helicobacter outer membrane proteins' (Hops), including adhesins BabA, SabA, HopQ, LabA and HopZ. Hops share a conserved 25 kDa C-terminal region that is thought to form an autotransporter-like transmembrane domain. Instead, our results show that Hops contain a non-continuous transmembrane domain, composed of seven predicted β-strands at the C-terminus and one at the N-terminus. Folding and outer membrane localization of the C-terminal β-domain critically depends on a predicted transmembrane β-strand within the first 16 N-terminal residues. The N-terminus is shown to reside in the periplasm, and our crystal and small angle X-ray scattering structures for the SabA extracellular domain reveal a conserved coiled-coil stem domain that connects to transmembrane β-strand 1 and 2. Taken together, our data show that Hop adhesins represent a novel outer membrane protein topology encompassing an OmpA-like 8-stranded β-barrel that is interrupted by a 15-108 kDa domain inserted inside the first extracellular loop. The insertion of large, folded domains in an extracellular loop is unprecedented in bacterial outer membrane proteins and is expected to have important consequences on how these proteins reach the cell surface.
- Structural and Molecular Microbiology, VIB-VUB Center for Structural Biology, VIB, Brussels, Belgium.
Organizational Affiliation: