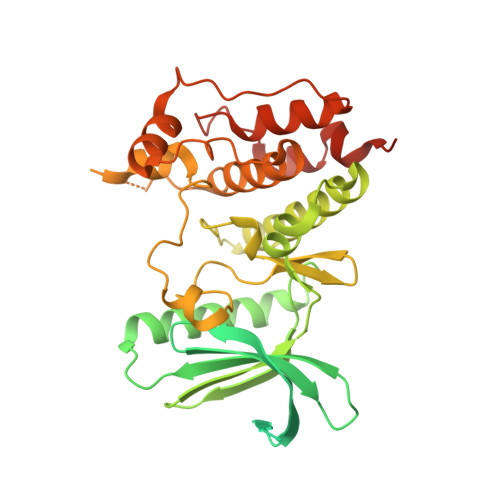
A crystal structure of the human protein kinase Mps1 reveals an ordered conformation of the activation loop.
Roorda, J.C., Joosten, R.P., Perrakis, A., Hiruma, Y.(2019) Proteins 87: 348-352
- PubMed: 30582207 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/prot.25651
- Primary Citation Related Structures:
6GVJ - PubMed Abstract:
Monopolar spindle 1 (Mps1) is a dual-specificity protein kinase, orchestrating faithful chromosome segregation during mitosis. All reported structures of the Mps1 kinase adopt the hallmarks of an inactive conformation, which includes a mostly disordered activation loop. Here, we present a 2.4 Å resolution crystal structure of an "extended" version of the Mps1 kinase domain, which shows an ordered activation loop. However, the other structural characteristics of an active kinase are not present. Our structure shows that the Mps1 activation loop can fit to the ATP binding pocket and interferes with ATP, but less so with inhibitors binding, partly explain the potency of various Mps1 inhibitors.
- Division of Biochemistry, Netherlands Cancer Institute, Amsterdam, The Netherlands.
Organizational Affiliation: