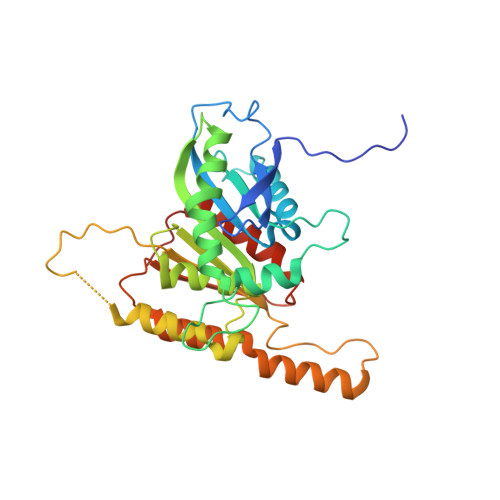
Iron Scavenging in Aspergillus Species: Structural and Biochemical Insights into Fungal Siderophore Esterases.
Ecker, F., Haas, H., Groll, M., Huber, E.M.(2018) Angew Chem Int Ed Engl 57: 14624-14629
- PubMed: 30070018
- DOI: https://doi.org/10.1002/anie.201807093
- Primary Citation Related Structures:
6GUD, 6GUG, 6GUI, 6GUL, 6GUN, 6GUO, 6GUP, 6GUR - PubMed Abstract:
Fungi utilize high-affinity chelators termed siderophores with chemically diverse structures to scavenge the essential nutrient iron from their surroundings. Since they are among the strongest known Fe 3+ binding agents, intracellular release of the heavy metal atom is facilitated by the activity of specific hydrolases. In this work, we report the characterization and X-ray crystal structures of four siderophore esterases: AfEstB and AfSidJ from Aspergillus fumigatus, as well as AnEstB and AnEstA from Aspergillus nidulans. Even though they all display the conserved α/β-hydrolase fold, we found significant structural and enzymatic discrepancies in their adaption to both related and chemically diverse substrates. A structure of AfEstB in complex with its substrate triacetylfusarinine C gives insight into the active enzyme and shows tetrahedral coordination between the catalytic serine and the scissile ester bond.
- Center for Integrated Protein Science Munich (CIPSM), Department of Chemistry, Technische Universität München, Lichtenbergstraße 4, 85748, Garching, Germany.
Organizational Affiliation: