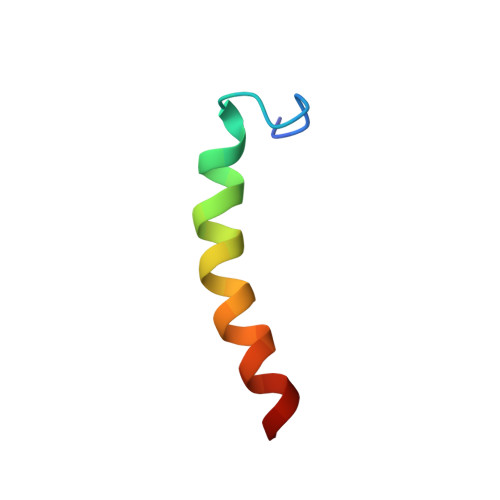
Structural insights into the AapA1 toxin of Helicobacter pylori.
Korkut, D.N., Alves, I.D., Vogel, A., Chabas, S., Sharma, C.M., Martinez, D., Loquet, A., Salgado, G.F., Darfeuille, F.(2020) Biochim Biophys Acta Gen Subj 1864: 129423-129423
- PubMed: 31476357 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbagen.2019.129423
- Primary Citation Related Structures:
6GIF, 6GIG - PubMed Abstract:
We previously reported the identification of the aapA1/IsoA1 locus as part of a new family of toxin-antitoxin (TA) systems in the human pathogen Helicobacter pylori. AapA1 belongs to type I TA bacterial toxins, and both its mechanism of action towards the membrane and toxicity features are still unclear. The biochemical characterization of the AapA1 toxic peptide was carried out using plasmid-borne expression and mutational approaches to follow its toxicity and localization. Biophysical properties of the AapA1 interaction with lipid membranes were studied by solution and solid-state NMR spectroscopy, plasmon waveguide resonance (PWR) and molecular modeling. We show that despite a low hydrophobic index, this toxin has a nanomolar affinity to the prokaryotic membrane. NMR spectroscopy reveals that the AapA1 toxin is structurally organized into three distinct domains: a positively charged disordered N-terminal domain (D), a single α-helix (H), and a basic C-terminal domain (R). The R domain interacts and destabilizes the membrane, while the H domain adopts a transmembrane conformation. These results were confirmed by alanine scanning of the minimal sequence required for toxicity. Our results have shown that specific amino acid residues along the H domain, as well as the R domain, are essential for the toxicity of the AapA1 toxin. Untangling and understanding the mechanism of action of small membrane-targeting toxins are difficult, but nevertheless contributes to a promising search and development of new antimicrobial drugs.
- INSERM, U1212, CNRS UMR 5320, Univ. Bordeaux, Laboratoire ARNA, F-33076 Bordeaux, France; Univ. Bordeaux, IECB, Laboratoire ARNA, F-33600 Pessac, France.
Organizational Affiliation: