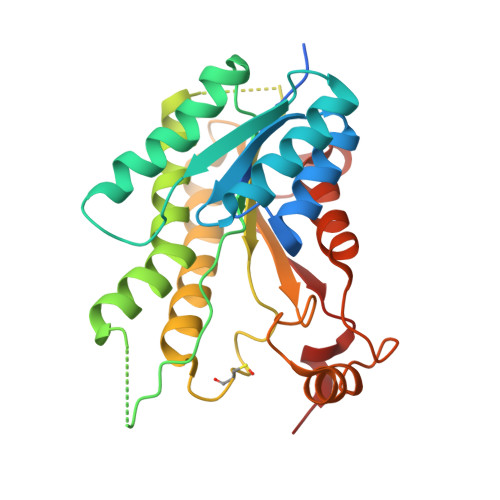
Enhancement of Benzothiazoles as Pteridine Reductase-1 Inhibitors for the Treatment of Trypanosomatidic Infections.
Linciano, P., Pozzi, C., Iacono, L.D., di Pisa, F., Landi, G., Bonucci, A., Gul, S., Kuzikov, M., Ellinger, B., Witt, G., Santarem, N., Baptista, C., Franco, C., Moraes, C.B., Muller, W., Wittig, U., Luciani, R., Sesenna, A., Quotadamo, A., Ferrari, S., Pohner, I., Cordeiro-da-Silva, A., Mangani, S., Costantino, L., Costi, M.P.(2019) J Med Chem 62: 3989-4012
- PubMed: 30908048 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.8b02021
- Primary Citation Related Structures:
6GCK, 6GCL, 6GCP, 6GCQ, 6GD0, 6GD4, 6GDO, 6GDP, 6GEX, 6GEY - PubMed Abstract:
2-Amino-benzo[ d]thiazole was identified as a new scaffold for the development of improved pteridine reductase-1 (PTR1) inhibitors and anti-trypanosomatidic agents. Molecular docking and crystallography guided the design and synthesis of 42 new benzothiazoles. The compounds were assessed for Trypanosoma brucei and Leishmania major PTR1 inhibition and in vitro activity against T. brucei and amastigote Leishmania infantum. We identified several 2-amino-benzo[ d]thiazoles with improved enzymatic activity ( TbPTR1 IC 50 = 0.35 μM; LmPTR1 IC 50 = 1.9 μM) and low μM antiparasitic activity against T. brucei. The ten most active compounds against TbPTR1 were able to potentiate the antiparasitic activity of methotrexate when evaluated in combination against T. brucei, with a potentiating index between 1.2 and 2.7. The compound library was profiled for early ADME toxicity, and 2-amino- N-benzylbenzo[ d]thiazole-6-carboxamide (4c) was finally identified as a novel potent, safe, and selective anti-trypanocydal agent (EC 50 = 7.0 μM). Formulation of 4c with hydroxypropyl-β-cyclodextrin yielded good oral bioavailability, encouraging progression to in vivo studies.
- Dipartimento di Scienze della Vita , University of Modena and Reggio Emilia , Via Campi 103 , 41125 Modena , Italy.
Organizational Affiliation: