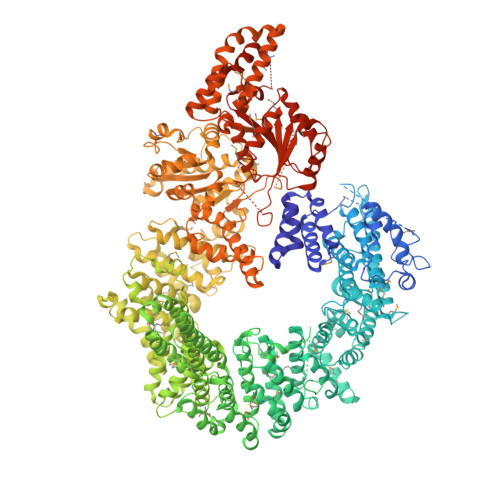
Crystal structure of the full Swi2/Snf2 remodeler Mot1 in the resting state.
Butryn, A., Woike, S., Shetty, S.J., Auble, D.T., Hopfner, K.P.(2018) Elife 7
- PubMed: 30289385
- DOI: https://doi.org/10.7554/eLife.37774
- Primary Citation Related Structures:
6G7E - PubMed Abstract:
Swi2/Snf2 ATPases remodel protein:DNA complexes in all of the fundamental chromosome-associated processes. The single-subunit remodeler Mot1 dissociates TATA box-binding protein (TBP):DNA complexes and provides a simple model for obtaining structural insights into the action of Swi2/Snf2 ATPases. Previously we reported how the N-terminal domain of Mot1 binds TBP, NC2 and DNA, but the location of the C-terminal ATPase domain remained unclear (Butryn et al., 2015). Here, we report the crystal structure of the near full-length Mot1 from Chaetomium thermophilum. Our data show that Mot1 adopts a ring like structure with a catalytically inactive resting state of the ATPase. Biochemical analysis suggests that TBP binding switches Mot1 into an ATP hydrolysis-competent conformation. Combined with our previous results, these data significantly improve the structural model for the complete Mot1:TBP:DNA complex and suggest a general mechanism for Mot1 action.
- Department of Biochemistry, Ludwig-Maximilians-Universität München, Munich, Germany.
Organizational Affiliation: