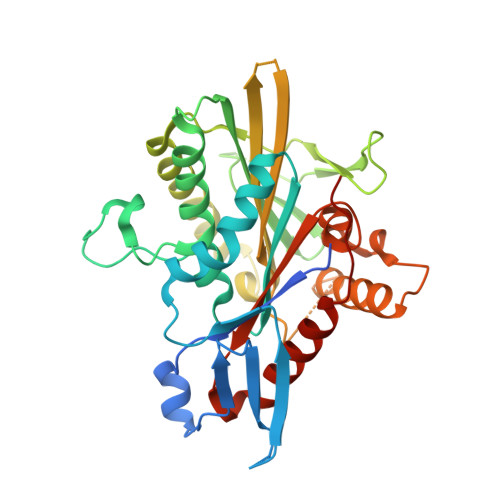
Crystal structure of the Eg5 - K858 complex and implications for structure-based design of thiadiazole-containing inhibitors.
Talapatra, S.K., Tham, C.L., Guglielmi, P., Cirilli, R., Chandrasekaran, B., Karpoormath, R., Carradori, S., Kozielski, F.(2018) Eur J Med Chem 156: 641-651
- PubMed: 30031975 Search on PubMed
- DOI: https://doi.org/10.1016/j.ejmech.2018.07.006
- Primary Citation Related Structures:
6G6Y, 6G6Z - PubMed Abstract:
The thiadiazole scaffold is an important core moiety in a variety of clinical drug candidates targeting a range of diseases. For example, the 2,4,5-substituted 1,3,4-thiadiazole scaffold is present in a lead compound and at least two clinical candidates targeting the human motor protein Eg5, against neoplastic diseases. An inhibitor named K858 has in vivo activity in various mouse xenografts whereas the clinical candidates (S)-ARRY-520 and (R)-Litronesib have entered clinical trials with the former one in phase III clinical trials either alone or in combination with a proteasome inhibitor against relapsed/refractory multiple myeloma. Astonishingly, structural data are lacking for all thiadiazole-containing Eg5 inhibitors. Here we report the structure determination of two crystal forms of the ternary Eg5-ADP-K858 complex, locking the motor in the so-called final inhibitor bound state, thus blocking ADP release, a crucial stage for Eg5 activity. K858 acts at the established allosteric inhibitor-binding pocket formed of helix α2, loop L5 and helix α3. The structure of the complex has far reaching consequences for thiadiazole containing Eg5 inhibitors. For example, we could rationalise the structure-activity relationship in the crucial 5-position of the thiadiazole scaffold and the complex will serve in the future as a basis for strucutre-based drug design.
- School of Pharmacy, University College London, 29-39 Brunswick Square, London, WC1N 1AX, United Kingdom.
Organizational Affiliation: