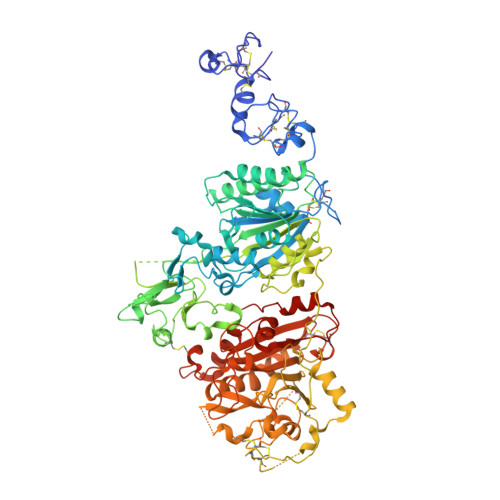
Crystallization of ectonucleotide phosphodiesterase/pyrophosphatase-3 and orientation of the SMB domains in the full-length ectodomain.
Dohler, C., Zebisch, M., Krinke, D., Robitzki, A., Strater, N.(2018) Acta Crystallogr F Struct Biol Commun 74: 696-703
- PubMed: 30387774 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X18011111
- Primary Citation Related Structures:
6G4G - PubMed Abstract:
Ectonucleotide phosphodiesterase/pyrophosphatase-3 (NPP3, ENPP3) is an ATP-hydrolyzing glycoprotein that is located in the extracellular space. The full-length ectodomain of rat NPP3 was expressed in HEK293S GntI - cells, purified using two chromatographic steps and crystallized. Its structure at 2.77 Å resolution reveals that the active-site zinc ions are missing and a large part of the active site and the surrounding residues are flexible. The SMB-like domains have the same orientation in all four molecules in the asymmetric unit. The SMB2 domain is oriented as in NPP2, but the SMB1 domain does not interact with the PDE domain but extends further away from the PDE domain. Deletion of the SMB domains resulted in crystals that diffracted to 2.4 Å resolution and are suitable for substrate-binding studies.
- Institute of Bioanalytical Chemistry, Center for Biotechnology and Biomedicine, Leipzig University, Deutscher Platz 5, 04103 Leipzig, Germany.
Organizational Affiliation: