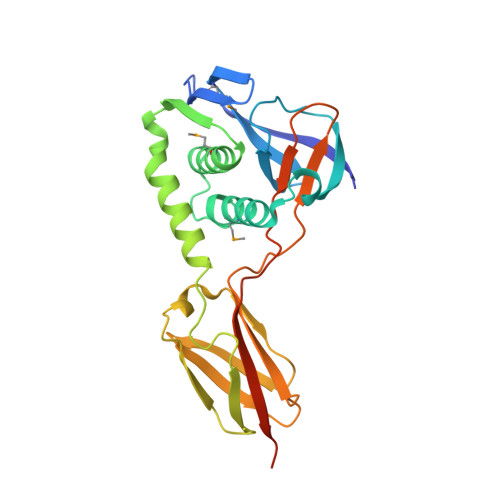
A new structural class of bacterial thioester domains reveals a slipknot topology.
Miller, O.K., Banfield, M.J., Schwarz-Linek, U.(2018) Protein Sci 27: 1651-1660
- PubMed: 30052296 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.3478
- Primary Citation Related Structures:
6FWV, 6FWY, 6FX6 - PubMed Abstract:
An increasing number of surface-associated proteins identified in Gram-positive bacteria are characterized by intramolecular cross-links in structurally conserved thioester, isopeptide, and ester domains (TIE proteins). Two classes of thioester domains (TEDs) have been predicted based on sequence with, to date, only representatives of Class I structurally characterized. Here, we present crystal structures of three Class II TEDs from Bacillus anthracis, vancomycin-resistant Staphylococcus aureus, and vancomycin-resistant Enterococcus faecium. These proteins are structurally distinct from Class I TEDs due to a β-sandwich domain that is inserted into the conserved TED fold to form a slipknot structure. Further, the B. anthracis TED domain is presented in the context of a full-length sortase-anchored protein structure (BaTIE). This provides insight into the three-dimensional arrangement of TIE proteins, which emerge as very abundant putative adhesins of Gram-positive bacteria.
- Biomedical Sciences Research Complex and School of Biology, University of St Andrews, St Andrews, KY16 9ST, United Kingdom.
Organizational Affiliation: