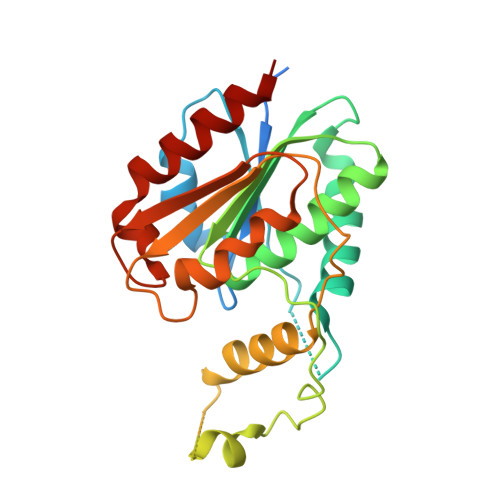
Biochemical and Structural Characterization of TesA, a Major Thioesterase Required for Outer-Envelope Lipid Biosynthesis in Mycobacterium tuberculosis.
Nguyen, P.C., Nguyen, V.S., Martin, B.P., Fourquet, P., Camoin, L., Spilling, C.D., Cavalier, J.F., Cambillau, C., Canaan, S.(2018) J Mol Biology 430: 5120-5136
- PubMed: 30292819 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2018.09.017
- Primary Citation Related Structures:
6FVJ, 6FW5 - PubMed Abstract:
With the high number of patients infected by tuberculosis and the sharp increase of drug-resistant tuberculosis cases, developing new drugs to fight this disease has become increasingly urgent. In this context, analogs of the naturally occurring enolphosphates Cyclipostins and Cyclophostin (CyC analogs) offer new therapeutic opportunities. The CyC analogs display potent activity both in vitro and in infected macrophages against several pathogenic mycobacteria including Mycobacterium tuberculosis and Mycobacterium abscessus. Interestingly, these CyC inhibitors target several enzymes with active-site serine or cysteine residues that play key roles in mycobacterial lipid and cell wall metabolism. Among them, TesA, a putative thioesterase involved in the synthesis of phthiocerol dimycocerosates (PDIMs) and phenolic glycolipids (PGLs), has been identified. These two lipids (PDIM and PGL) are non-covalently bound to the outer cell wall in several human pathogenic mycobacteria and are important virulence factors. Herein, we used biochemical and structural approaches to validate TesA as an effective pharmacological target of the CyC analogs. We confirmed both thioesterase and esterase activities of TesA, and showed that the most active inhibitor CyC 17 binds covalently to the catalytic Ser104 residue leading to a total loss of enzyme activity. These data were supported by the X-ray structure, obtained at a 2.6-Å resolution, of a complex in which CyC 17 is bound to TesA. Our study provides evidence that CyC 17 inhibits the activity of TesA, thus paving the way to a new strategy for impairing the PDIM and PGL biosynthesis, potentially decreasing the virulence of associated mycobacterial species.
- Aix-Marseille University, CNRS, LISM, IMM FR3479, Marseille, France.
Organizational Affiliation: