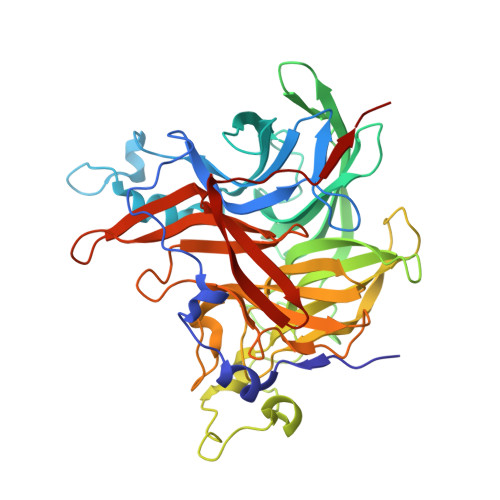
Comparison of the Levansucrase from the epiphyte Erwinia tasmaniensis vs its homologue from the phytopathogen Erwinia amylovora.
Polsinelli, I., Caliandro, R., Salomone-Stagni, M., Demitri, N., Rejzek, M., Field, R.A., Benini, S.(2019) Int J Biol Macromol 127: 496-501
- PubMed: 30660564
- DOI: https://doi.org/10.1016/j.ijbiomac.2019.01.074
- Primary Citation Related Structures:
6FRW - PubMed Abstract:
Erwinia tasmaniensis is an epiphytic bacterium related to the plant pathogen Erwinia amylovora, the etiological agent of fire blight. In this study the levansucrase from E. tasmaniensis (EtLsc) has been compared with the homologous enzyme from E. amylovora (EaLsc). We characterized the enzymatic activity and compared the products profile of both enzymes by High Performance Anion Exchange Chromatography coupled with Pulsed Amperometric Detector (HPAEC-PAD). Moreover we determined the crystal structure of EtLsc to understand the structural peculiarity causing the different product profiles of the two homologues. EtLsc exhibits increased efficiency in the production of FOS, resulting in a better catalyst for biotechnological synthesis than EaLsc. Based on our results, we propose that the role of this enzyme in the life cycle of the two bacteria is most likely related to survival, rather than linked to pathogenicity in E. amylovora.
- Bioorganic Chemistry and Bio-Crystallography laboratory (B(2)Cl), Faculty of Science and Technology, Free University of Bolzano, Piazza Università 5, 39100 Bolzano, Italy.
Organizational Affiliation: