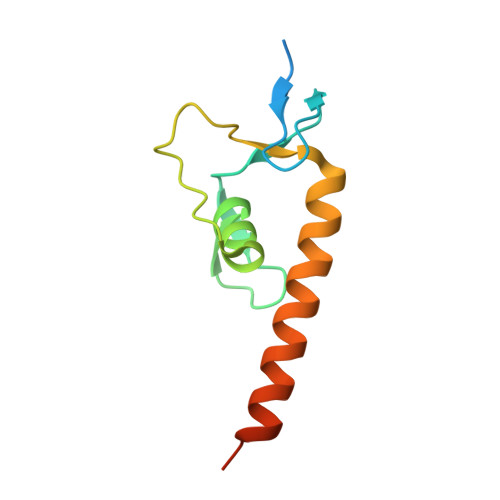
Structural basis of diversity and homodimerization specificity of zinc-finger-associated domains in Drosophila.
Bonchuk, A., Boyko, K., Fedotova, A., Nikolaeva, A., Lushchekina, S., Khrustaleva, A., Popov, V., Georgiev, P.(2021) Nucleic Acids Res 49: 2375-2389
- PubMed: 33638995
- DOI: https://doi.org/10.1093/nar/gkab061
- Primary Citation Related Structures:
6FP5 - PubMed Abstract:
In arthropods, zinc finger-associated domains (ZADs) are found at the N-termini of many DNA-binding proteins with tandem arrays of Cys2-His2 zinc fingers (ZAD-C2H2 proteins). ZAD-C2H2 proteins undergo fast evolutionary lineage-specific expansion and functional diversification. Here, we show that all ZADs from Drosophila melanogaster form homodimers, but only certain ZADs with high homology can also heterodimerize. CG2712, for example, is unable to heterodimerize with its paralog, the previously characterized insulator protein Zw5, with which it shares 46% homology. We obtained a crystal structure of CG2712 protein's ZAD domain that, in spite of a low sequence homology, has similar spatial organization with the only known ZAD structure (from Grauzone protein). Steric clashes prevented the formation of heterodimers between Grauzone and CG2712 ZADs. Using detailed structural analysis, site-directed mutagenesis, and molecular dynamics simulations, we demonstrated that rapid evolutionary acquisition of interaction specificity was mediated by the more energy-favorable formation of homodimers in comparison to heterodimers, and that this specificity was achieved by multiple amino acid substitutions resulting in the formation or breaking of stabilizing interactions. We speculate that specific homodimerization of ZAD-C2H2 proteins is important for their architectural role in genome organization.
- Department of the Control of Genetic Processes, Institute of Gene Biology, Russian Academy of Sciences, Moscow 119334, Russia.
Organizational Affiliation: