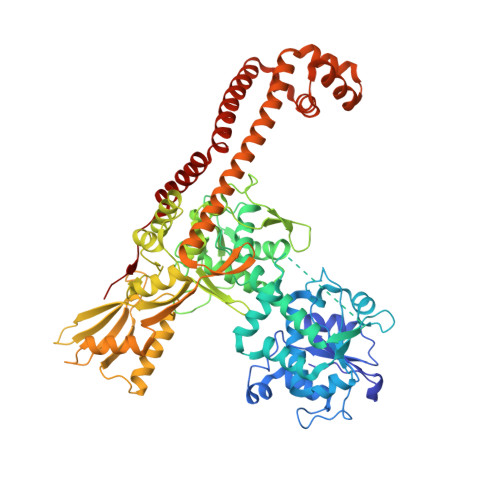
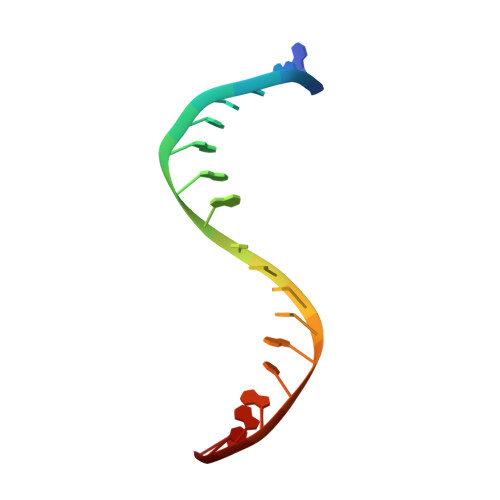
Virtual Screening Approach and Investigation of Structure-Activity Relationships To Discover Novel Bacterial Topoisomerase Inhibitors Targeting Gram-Positive and Gram-Negative Pathogens.
Magaro, G., Prati, F., Garofalo, B., Corso, G., Furlotti, G., Apicella, C., Mangano, G., D'Atanasio, N., Robinson, D., Di Giorgio, F.P., Ombrato, R.(2019) J Med Chem 62: 7445-7472
- PubMed: 31276392 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.9b00394
- Primary Citation Related Structures:
6FM4 - PubMed Abstract:
Bacterial resistance is increasing rapidly, requiring urgent identification of new antibacterial drugs that are effective against multidrug-resistant pathogens. Novel bacterial topoisomerase inhibitors (NBTIs) provide a new strategy for investigating the well-validated DNA gyrase and topoisomerase IV targets while preventing cross-resistance issues. On this basis, starting from a virtual screening campaign and subsequent structure-based hit optimization guided by X-ray studies, a novel class of piperazine-like NBTIs with outstanding enzymatic activity against Staphylococcus aureus and Escherichia coli DNA gyrase and topoisomerase IV was identified. Notably, compounds (±)-33 , (±)-35 , and (±)-36 with potent and balanced multitarget enzymatic profiles exhibited excellent efficacy against selected Gram-positive and Gram-negative pathogens, as well as clinically relevant resistant strains. Overall, the new NBTI chemotype described herein, owing to the broad-spectrum antibacterial activity and favorable in vitro safety profile, might serve as a basis for the development of novel treatments against serious infections.
- Angelini RR&D (Research, Regulatory & Development) , Angelini S.p.A. , Piazzale della Stazione SNC, S. Palomba-Pomezia , Rome 00071 , Italy.
Organizational Affiliation: