High-affinity ligands of the colchicine domain in tubulin based on a structure-guided design.
Bueno, O., Estevez Gallego, J., Martins, S., Prota, A.E., Gago, F., Gomez-SanJuan, A., Camarasa, M.J., Barasoain, I., Steinmetz, M.O., Diaz, J.F., Perez-Perez, M.J., Liekens, S., Priego, E.M.(2018) Sci Rep 8: 4242-4242
- PubMed: 29523799 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-018-22382-x
- Primary Citation Related Structures:
6FKJ, 6FKL - PubMed Abstract:
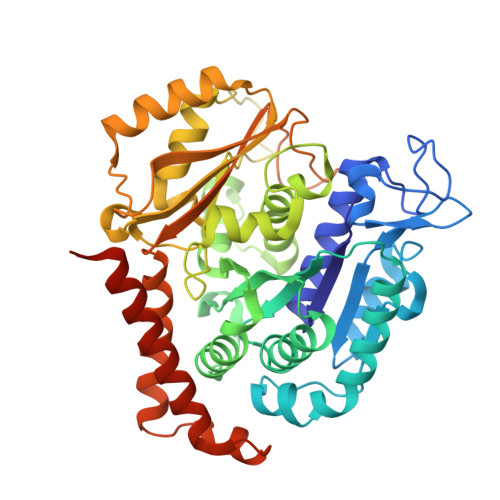
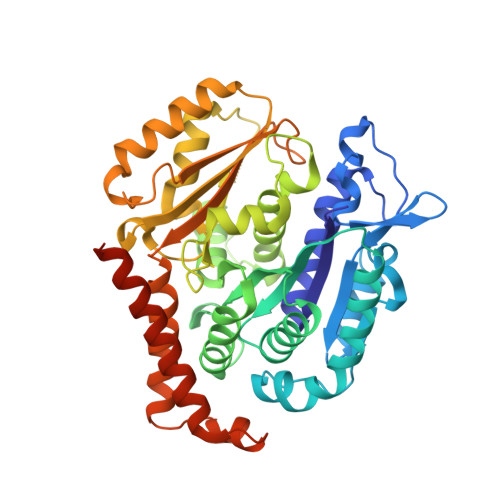
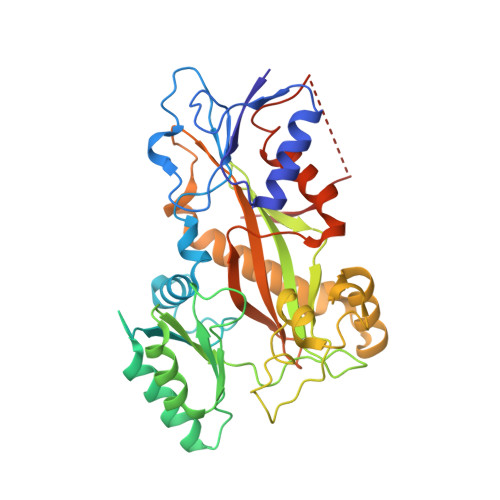
Microtubule-targeting agents that bind at the colchicine-site of tubulin are of particular interest in antitumoral therapy due to their dual mechanism of action as antimitotics and vascular disrupting agents. Cyclohexanediones derivatives have been described as a new family of colchicine-domain binders with an association constant to tubulin similar to that of colchicine. Here, the high-resolution structures of tubulin in complex with cyclohexanediones TUB015 and TUB075 were solved by X-ray crystallography. A detailed analysis of the tubulin-TUB075 interaction by means of computational affinity maps allowed the identification of two additional regions at the binding site that were addressed with the design and synthesis of a new series of cyclohexanediones with a distal 2-substituted benzofurane. These new compounds showed potent antiproliferative activity with IC 50 values in the nM range, arrested cell cycle progression at the G 2 /M phase and induced apoptosis at sub μM concentrations. Moreover, they caused the destruction of a preformed vascular network in vitro and inhibited the migration of endothelial cells at non-toxic concentrations. Finally, these compounds displayed high affinity for tubulin as substantiated by a K b value of 2.87 × 10 8 M -1 which, to the best of our knowledge, represents the highest binding constant measured to date for a colchicine-domain ligand.
- Instituto de Química Médica (IQM,CSIC), Juan de la Cierva 3, 28006, Madrid, Spain.
Organizational Affiliation: