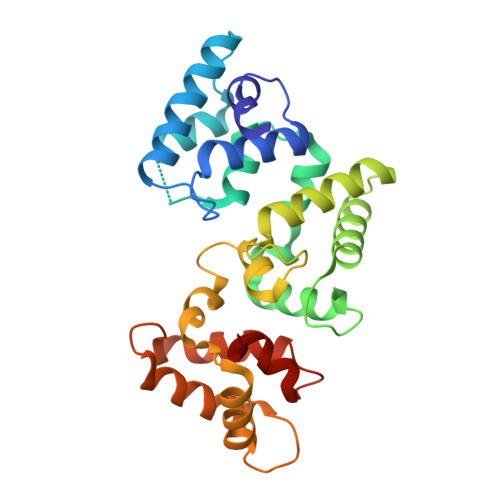
The X-ray structure of human calbindin-D28K: an improved model.
Noble, J.W., Almalki, R., Roe, S.M., Wagner, A., Duman, R., Atack, J.R.(2018) Acta Crystallogr D Struct Biol 74: 1008-1014
- PubMed: 30289411 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2059798318011610
- Primary Citation Related Structures:
6FIE - PubMed Abstract:
Calbindin-D28K is a widely expressed calcium-buffering cytoplasmic protein that is involved in many physiological processes. It has been shown to interact with other proteins, suggesting a role as a calcium sensor. Many of the targets of calbindin-D28K are of therapeutic interest: for example, inositol monophosphatase, the putative target of lithium therapy in bipolar disorder. Presented here is the first crystal structure of human calbindin-D28K. There are significant deviations in the tertiary structure when compared with the NMR structure of rat calbindin-D28K (PDB entry 2g9b), despite 98% sequence identity. Small-angle X-ray scattering (SAXS) indicates that the crystal structure better predicts the properties of calbindin-D28K in solution compared with the NMR structure. Here, the first direct visualization of the calcium-binding properties of calbindin-D28K is presented. Four of the six EF-hands that make up the secondary structure of the protein contain a calcium-binding site. Two distinct conformations of the N-terminal EF-hand calcium-binding site were identified using long-wavelength calcium single-wavelength anomalous dispersion (SAD). This flexible region has previously been recognized as a protein-protein interaction interface. SAXS data collected in both the presence and absence of calcium indicate that there are no large structural differences in the globular structure of calbindin-D28K between the calcium-loaded and unloaded proteins.
- Sussex Drug Discovery Centre, University Of Sussex, Falmer, Brighton BN1 9QG, England.
Organizational Affiliation: