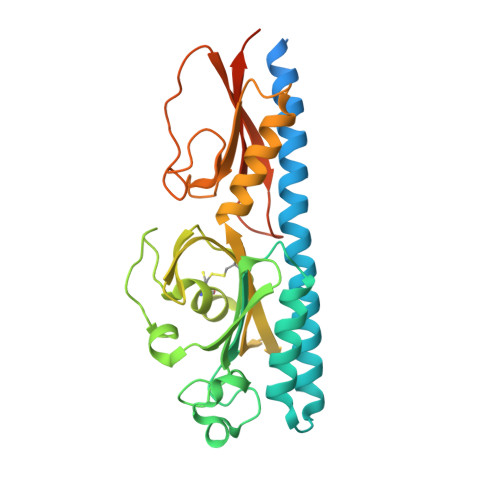
Structural Basis for Polyamine Binding at the dCACHE Domain of the McpU Chemoreceptor from Pseudomonas putida.
Gavira, J.A., Ortega, A., Martin-Mora, D., Conejero-Muriel, M.T., Corral-Lugo, A., Morel, B., Matilla, M.A., Krell, T.(2018) J Mol Biology 430: 1950-1963
- PubMed: 29758259 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2018.05.008
- Primary Citation Related Structures:
6F9G - PubMed Abstract:
Many bacteria can move chemotactically to a variety of compounds and the recognition of chemoeffectors by the chemoreceptor ligand binding domain (LBD) defines the specificity of response. Many chemoreceptors were found to recognize different amino and organic acids, but the McpU chemoreceptor from Pseudomonas putida was identified as the first chemoreceptor that bound specifically polyamines. We report here the three-dimensional structure of McpU-LBD in complex with putrescine at a resolution of 2.4 Å, which fitted well a solution structure generated by small-angle X-ray scattering. Putrescine bound to a negatively charged pocket in the membrane distal module of McpU-LBD. Similarities exist in the binding of putrescine to McpU-LBD and taurine to the LBD of the Mlp37 chemoreceptor of Vibrio cholerae. In both structures, the primary amino group of the respective ligand is recognized by hydrogen bonds established by two aspartate and a tyrosine side chain. This feature may be used to predict the ligands of chemoreceptors with unknown function. Analytical ultracentrifugation revealed that McpU-LBD is monomeric in solution and that ligand binding does not alter this oligomeric state. This sensing mode thus differs from that of the well-characterised four-helix bundle domains where ligands bind to two sites at the LBD dimer interface. Although there appear to be different sensing modes, results are discussed in the context of data, indicating that chemoreceptors employ the same mechanism of transmembrane signaling. This work enhances our understanding of CACHE domains, which are the most abundant sensor domains in bacterial chemoreceptors and sensor kinases.
- Laboratory of Crystallographic Studies, IACT, (CSIC-UGR), Avenida de las Palmeras 4, 18100 Armilla, Granada, Spain.
Organizational Affiliation: