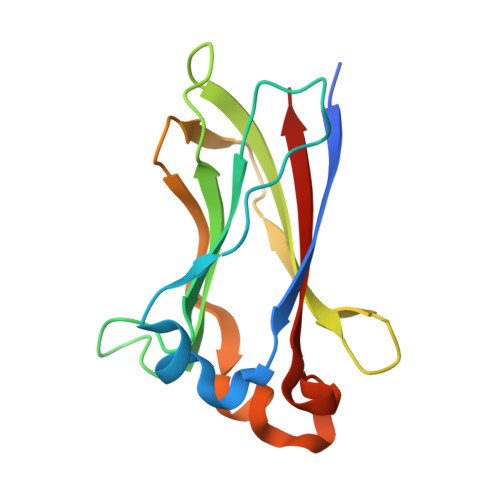
The Structure of the SPOP-Pdx1 Interface Reveals Insights into the Phosphorylation-Dependent Binding Regulation.
Ostertag, M.S., Messias, A.C., Sattler, M., Popowicz, G.M.(2019) Structure 27: 327-334.e3
- PubMed: 30449689 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2018.10.005
- Primary Citation Related Structures:
6F8F, 6F8G - PubMed Abstract:
Pdx1 is a transcription factor crucial for development and maintenance of a functional pancreas. It regulates insulin expression and glucose homeostasis. SPOP is an E3-ubiquitin ligase adaptor protein that binds Pdx1, thus triggering its ubiquitination and proteasomal degradation. However, the underlying mechanisms are not well understood. Here, we present the crystal structure of the SPOP-Pdx1 complex. We show that Pdx1 residues 223-233 bind to SPOP MATH domain with low micromolar affinity. The interface is extended compared to other SPOP-client proteins. Previously, Pdx1 phosphorylation has been proposed to have a regulatory function. In this respect we show that phosphorylation lowers the affinity of Pdx1 to SPOP by isothermal titration calorimetry and nuclear magnetic resonance data. Our data provide insights into a critical protein-protein interaction that regulates cellular Pdx1 levels by SPOP-mediated decay. A reduction of Pdx1 levels in β cells is linked to apoptosis and considered a hallmark of type 2 diabetes.
- Institute of Structural Biology, Helmholtz Zentrum München, Ingolstädter Landstr. 1, Neuherberg 85764, Germany; Center for Integrated Protein Science Munich at Chair of Biomolecular NMR Spectroscopy, Department Chemie, Technische Universität München, Lichtenbergstrasse 4, Garching 85747, Germany.
Organizational Affiliation: