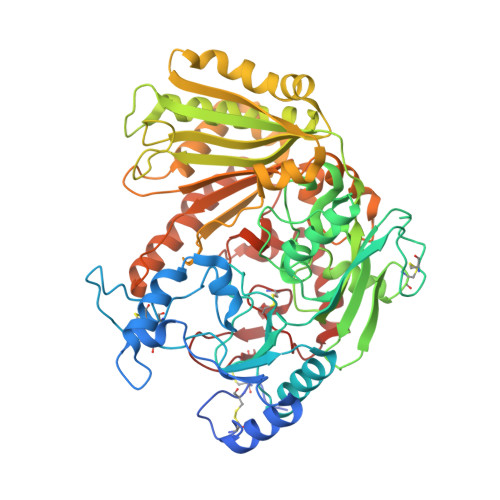
Characterization of Two VAO-Type Flavoprotein Oxidases from Myceliophthora thermophila.
Ferrari, A.R., Rozeboom, H.J., Vugts, A.S.C., Koetsier, M.J., Floor, R., Fraaije, M.W.(2018) Molecules 23
- PubMed: 29303991 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/molecules23010111
- Primary Citation Related Structures:
6F72, 6F73, 6F74 - PubMed Abstract:
The VAO flavoprotein family consists mostly of oxidoreductases harboring a covalently linked flavin cofactor. The linkage can be either monocovalent at position 8 with a histidine or tyrosine or bicovalent at position 8 with a histidine and at position 6 with a cysteine. Bicovalently bound flavoproteins show a preference for bulkier substrates such as oligosaccharides or secondary metabolites. The genome of the thermophilic fungus Myceliophthora thermophila C1 was found to be rich in genes encoding putative covalent VAO-type flavoproteins. Enzymes from this fungus have the advantage of being rather thermostable and homologous overexpression in M. thermophila C1 is feasible. Recently we discovered a new and VAO-type carbohydrate oxidase from this fungus: xylooligosaccharide oxidase. In this study, two other putative VAO-type oxidases, protein sequence XP_003663615 (MtVAO615) and XP_003665713 (MtVAO713), were expressed in M. thermophila C1, purified and characterized. Enzyme MtVAO615 was found to contain a bicovalently bound FAD, while enzyme MtVAO713 contained a monocovalent histidyl-bound FAD. The crystal structures of both proteins were obtained which revealed atypical active site architectures. It could be experimentally verified that both proteins, when reduced, rapidly react with molecular oxygen, a hallmark of flavoprotein oxidases. A large panel of alcohols, including carbohydrates, steroids and secondary alcohols were tested as potential substrates. For enzyme MtVAO713 low oxidase activity was discovered towards ricinoleic acid.
- Molecular Enzymology, Groningen Biomolecular Sciences and Biotechnology Institute, University of Groningen, 9747AG Groningen, The Netherlands. a.ferrari88@gmail.com.
Organizational Affiliation: