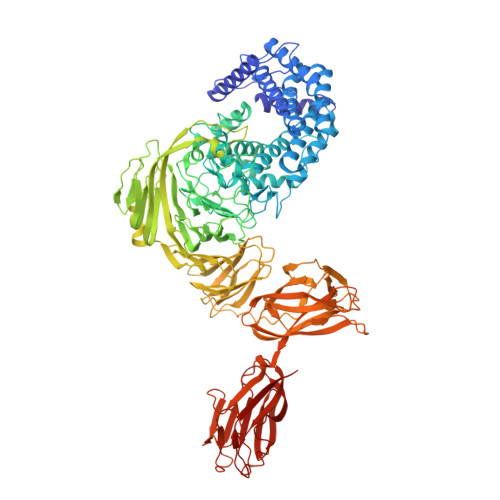
Structure and Dynamics of a Promiscuous Xanthan Lyase from Paenibacillus nanensis and the Design of Variants with Increased Stability and Activity.
Jensen, P.F., Kadziola, A., Comamala, G., Segura, D.R., Anderson, L., Poulsen, J.N., Rasmussen, K.K., Agarwal, S., Sainathan, R.K., Monrad, R.N., Svendsen, A., Nielsen, J.E., Lo Leggio, L., Rand, K.D.(2019) Cell Chem Biol 26: 191-202.e6
- PubMed: 30503284 Search on PubMed
- DOI: https://doi.org/10.1016/j.chembiol.2018.10.016
- Primary Citation Related Structures:
6F2P - PubMed Abstract:
We have characterized the structure and dynamics of the carbohydrate-modifying enzyme Paenibacillus nanensis xanthan lyase (PXL) involved in the degradation of xanthan by X-ray crystallography, small-angle X-ray scattering, and hydrogen/deuterium exchange mass spectrometry. Unlike other xanthan lyases, PXL is specific for both unmodified mannose and pyruvylated mannose, which we find is correlated with structural differences in the substrate binding groove. The structure of the full-length enzyme reveals two additional C-terminal modules, one of which belongs to a new non-catalytic carbohydrate binding module family. Ca 2+ are critical for the activity and conformation of PXL, and we show that their removal by chelating agents results in localized destabilization/unfolding of particularly the C-terminal modules. We use the structure and the revealed impact of Ca 2+ coordination on conformational dynamics to guide the engineering of PXL variants with increased activity and stability in a chelating environment, thus expanding the possibilities for industrial applications of PXL.
- Protein Analysis Group, Department of Pharmacy, University of Copenhagen, Copenhagen 2100, Denmark.
Organizational Affiliation: