Born to sense: biophysical analyses of the oxygen sensing prolyl hydroxylase from the simplest animal Trichoplax adhaerens.
Lippl, K., Boleininger, A., McDonough, M.A., Abboud, M.I., Tarhonskaya, H., Chowdhury, R., Loenarz, C., Schofield, C.J.(2018) Hypoxia (Auckl) 6: 57-71
- PubMed: 30519597 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.2147/HP.S174655
- Primary Citation Related Structures:
6EY1, 6F0W - PubMed Abstract:
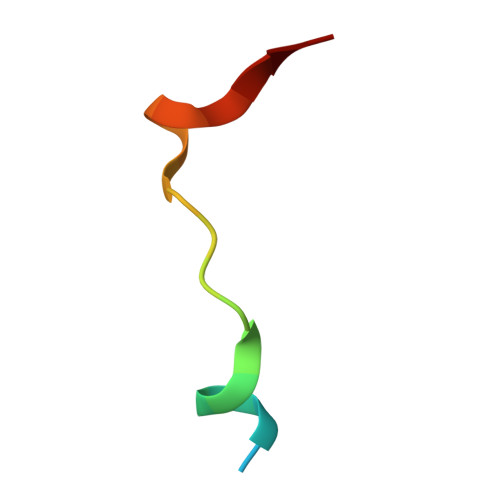
In humans and other animals, the chronic hypoxic response is mediated by hypoxia inducible transcription factors (HIFs) which regulate the expression of genes that counteract the effects of limiting oxygen. Prolyl hydroxylases (PHDs) act as hypoxia sensors for the HIF system in organisms ranging from humans to the simplest animal Trichoplax adhaerens . We report structural and biochemical studies on the T. adhaerens HIF prolyl hydroxylase ( Ta PHD) that inform about the evolution of hypoxia sensing in animals. High resolution crystal structures (≤1.3 Å) of Ta PHD, with and without its HIFα substrate, reveal remarkable conservation of key active site elements between T. adhaerens and human PHDs, which also manifest in kinetic comparisons. Conserved structural features of Ta PHD and human PHDs include those apparently enabling the slow binding/reaction of oxygen with the active site Fe(II), the formation of a stable 2-oxoglutarate complex, and a stereoelectronically promoted change in conformation of the hydroxylated proline-residue. Comparison of substrate selectivity between the human PHDs and Ta PHD provides insights into the selectivity determinants of HIF binding by the PHDs, and into the evolution of the multiple HIFs and PHDs present in higher animals.
- Chemistry Research Laboratory, University of Oxford, Oxford, UK, christopher.schofield@chem.ox.ac.uk.
Organizational Affiliation: