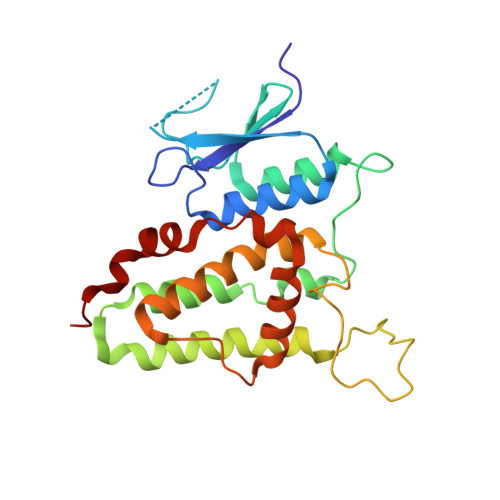
Inherent flexibility of CLIC6 revealed by crystallographic and solution studies.
Ferofontov, A., Strulovich, R., Marom, M., Giladi, M., Haitin, Y.(2018) Sci Rep 8: 6882-6882
- PubMed: 29720717 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-018-25231-z
- Primary Citation Related Structures:
6ERY, 6ERZ - PubMed Abstract:
Chloride intracellular channels (CLICs) are a family of unique proteins, that were suggested to adopt both soluble and membrane-associated forms. Moreover, following this unusual metamorphic change, CLICs were shown to incorporate into membranes and mediate ion conduction in vitro, suggesting multimerization upon membrane insertion. Here, we present a 1.8 Å resolution crystal structure of the CLIC domain of mouse CLIC6 (mCLIC6). The structure reveals a monomeric arrangement and shows a high degree of structural conservation with other CLICs. Small-angle X-ray scattering (SAXS) analysis of mCLIC6 demonstrated that the overall solution structure is similar to the crystallographic conformation. Strikingly, further analysis of the SAXS data using ensemble optimization method unveiled additional elongated conformations, elucidating high structural plasticity as an inherent property of the protein. Moreover, structure-guided perturbation of the inter-domain interface by mutagenesis resulted in a population shift towards elongated conformations of mCLIC6. Additionally, we demonstrate that oxidative conditions induce an increase in mCLIC6 hydrophobicity along with mild oligomerization, which was enhanced by the presence of membrane mimetics. Together, these results provide mechanistic insights into the metamorphic nature of mCLIC6.
- Department of Physiology and Pharmacology, Sackler School of Medicine, Tel-Aviv University, Tel-Aviv, 6997801, Israel.
Organizational Affiliation: