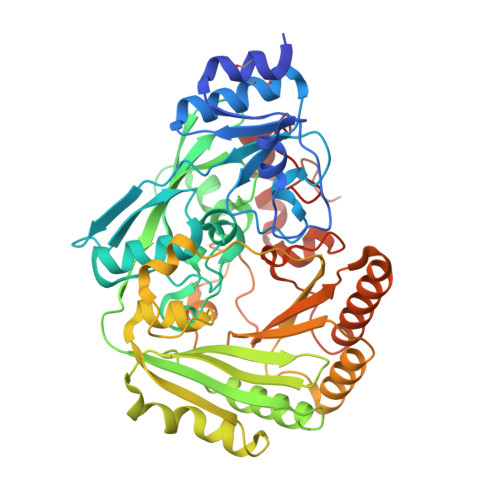
The single berberine bridge enzyme homolog of Physcomitrella patens is a cellobiose oxidase.
Toplak, M., Wiedemann, G., Ulicevic, J., Daniel, B., Hoernstein, S.N.W., Kothe, J., Niederhauser, J., Reski, R., Winkler, A., Macheroux, P.(2018) FEBS J 285: 1923-1943
- PubMed: 29633551
- DOI: https://doi.org/10.1111/febs.14458
- Primary Citation Related Structures:
6EO4, 6EO5 - PubMed Abstract:
The berberine bridge enzyme from the California poppy Eschscholzia californica (EcBBE) catalyzes the oxidative cyclization of (S)-reticuline to (S)-scoulerine, that is, the formation of the berberine bridge in the biosynthesis of benzylisoquinoline alkaloids. Interestingly, a large number of BBE-like genes have been identified in plants that lack alkaloid biosynthesis. This finding raised the question of the primordial role of BBE in the plant kingdom, which prompted us to investigate the closest relative of EcBBE in Physcomitrella patens (PpBBE1), the most basal plant harboring a BBE-like gene. Here, we report the biochemical, structural, and in vivo characterization of PpBBE1. Our studies revealed that PpBBE1 is structurally and biochemically very similar to EcBBE. In contrast to EcBBE, we found that PpBBE1 catalyzes the oxidation of the disaccharide cellobiose to the corresponding lactone, that is, PpBBE1 is a cellobiose oxidase. The enzymatic reaction mechanism was characterized by a structure-guided mutagenesis approach that enabled us to assign a catalytic role to amino acid residues in the active site of PpBBE1. In vivo experiments revealed the highest level of PpBBE1 expression in chloronema, the earliest stage of the plant's life cycle, where carbon metabolism is strongly upregulated. It was also shown that the enzyme is secreted to the extracellular space, where it may be involved in later steps of cellulose degradation, thereby allowing the moss to make use of cellulose for energy production. Overall, our results suggest that the primordial role of BBE-like enzymes in plants revolved around primary metabolic reactions in carbohydrate utilization. Structural data are available in the PDB under the accession numbers 6EO4 and 6EO5.
- Institute of Biochemistry, Graz University of Technology, Austria.
Organizational Affiliation: