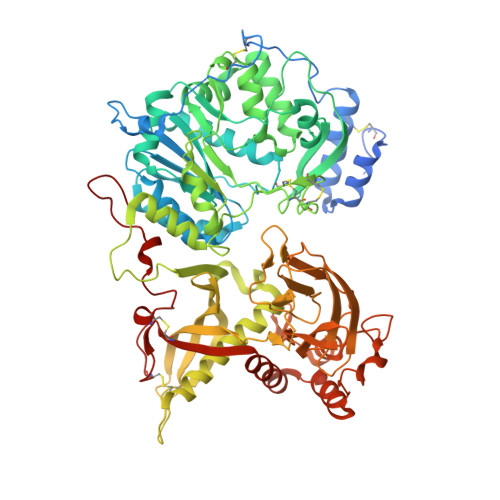
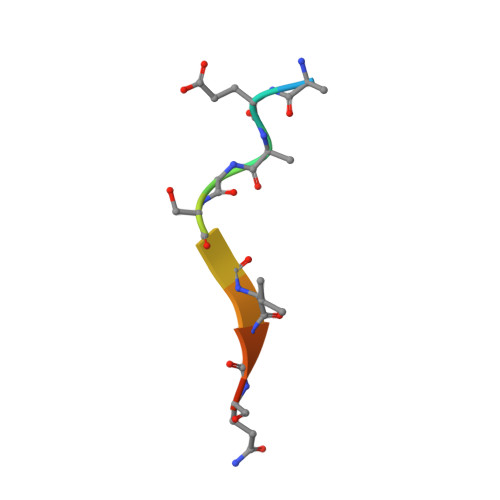
Structural Basis for the Initiation of Glycosaminoglycan Biosynthesis by Human Xylosyltransferase 1.
Briggs, D.C., Hohenester, E.(2018) Structure 26: 801-809.e3
- PubMed: 29681470 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2018.03.014
- Primary Citation Related Structures:
6EJ7, 6EJ8, 6EJ9, 6EJA, 6EJB, 6EJC, 6EJD, 6EJE, 6FOA - PubMed Abstract:
Proteoglycans (PGs) are essential components of the animal extracellular matrix and are required for cell adhesion, migration, signaling, and immune function. PGs are composed of a core protein and long glycosaminoglycan (GAG) chains, which often specify PG function. GAG biosynthesis is initiated by peptide O-xylosyltransferases, which transfer xylose onto selected serine residues in the core proteins. We have determined crystal structures of human xylosyltransferase 1 (XT1) in complex with the sugar donor, UDP-xylose, and various acceptor peptides. The structures reveal unique active-site features that, in conjunction with functional experiments, explain the substrate specificity of XT1. A constriction within the peptide binding cleft requires the acceptor serine to be followed by glycine or alanine. The remainder of the cleft can accommodate a wide variety of sequences, but with a general preference for acidic residues. These findings provide a framework for understanding the selectivity of GAG attachment.
- Department of Life Sciences, Imperial College London, London SW7 2AZ, UK. Electronic address: david.briggs@crick.ac.uk.
Organizational Affiliation: