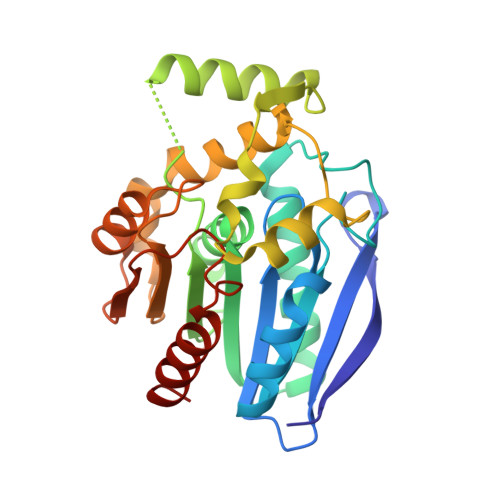
The crystal structure of monoacylglycerol lipase from M. tuberculosis reveals the basis for specific inhibition.
Aschauer, P., Zimmermann, R., Breinbauer, R., Pavkov-Keller, T., Oberer, M.(2018) Sci Rep 8: 8948-8948
- PubMed: 29895832
- DOI: https://doi.org/10.1038/s41598-018-27051-7
- Primary Citation Related Structures:
6EIC - PubMed Abstract:
Monoacylglycerol lipases (MGLs) are enzymes that hydrolyze monoacylglycerol into a free fatty acid and glycerol. Fatty acids can be used for triacylglycerol synthesis, as energy source, as building blocks for energy storage, and as precursor for membrane phospholipids. In Mycobacterium tuberculosis, fatty acids also serve as precursor for polyketide lipids like mycolic acids, major components of the cellular envelope associated to resistance for drug. We present the crystal structure of the MGL Rv0183 from Mycobacterium tuberculosis (mtbMGL) in open conformation. The structure reveals remarkable similarities with MGL from humans (hMGL) in both, the cap region and the α/β core. Nevertheless, mtbMGL could not be inhibited with JZL-184, a known inhibitor of hMGL. Docking studies provide an explanation why the activity of mtbMGL was not affected by the inhibitor. Our findings suggest that specific inhibition of mtbMGL from Mycobacterium tuberculosis, one of the oldest recognized pathogens, is possible without influencing hMGL.
- Institute of Molecular Biosciences, University of Graz, Humboldtstraße 50/3, 8010, Graz, Austria.
Organizational Affiliation: