Antibodies to a Conserved Influenza Head Interface Epitope Protect by an IgG Subtype-Dependent Mechanism.
Watanabe, A., McCarthy, K.R., Kuraoka, M., Schmidt, A.G., Adachi, Y., Onodera, T., Tonouchi, K., Caradonna, T.M., Bajic, G., Song, S., McGee, C.E., Sempowski, G.D., Feng, F., Urick, P., Kepler, T.B., Takahashi, Y., Harrison, S.C., Kelsoe, G.(2019) Cell 177: 1124-1135.e16
- PubMed: 31100267 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2019.03.048
- Primary Citation Related Structures:
6E4X, 6E56 - PubMed Abstract:
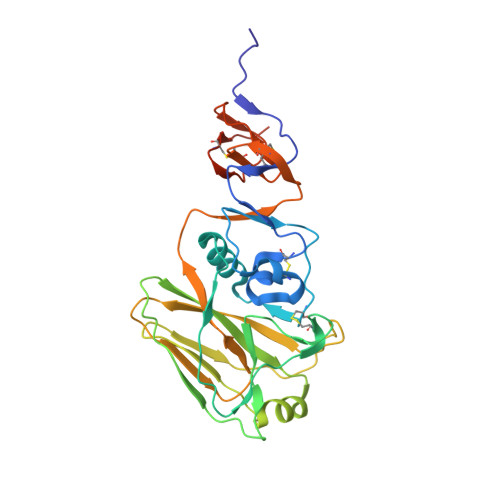
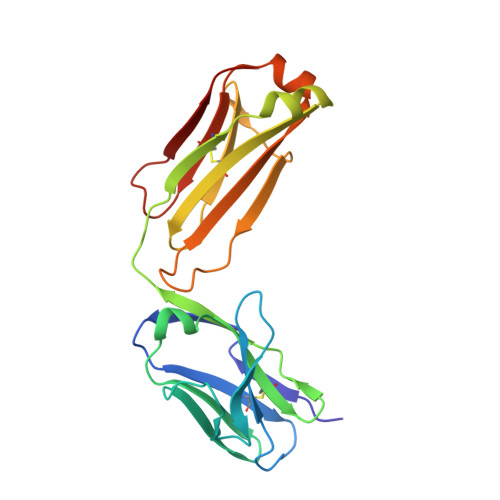
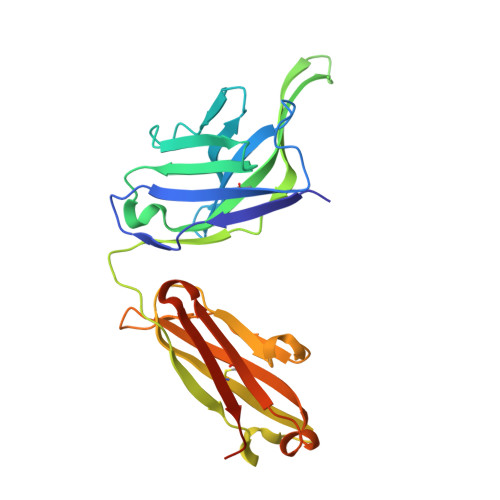
Vaccines to generate durable humoral immunity against antigenically evolving pathogens such as the influenza virus must elicit antibodies that recognize conserved epitopes. Analysis of single memory B cells from immunized human donors has led us to characterize a previously unrecognized epitope of influenza hemagglutinin (HA) that is immunogenic in humans and conserved among influenza subtypes. Structures show that an unrelated antibody from a participant in an experimental infection protocol recognized the epitope as well. IgGs specific for this antigenic determinant do not block viral infection in vitro, but passive administration to mice affords robust IgG subtype-dependent protection against influenza infection. The epitope, occluded in the pre-fusion form of HA, is at the contact surface between HA head domains; reversible molecular "breathing" of the HA trimer can expose the interface to antibody and B cells. Antigens that present this broadly immunogenic HA epitope may be good candidates for inclusion in "universal" flu vaccines.
- Department of Immunology, Duke University, Durham, NC 27710, USA.
Organizational Affiliation: