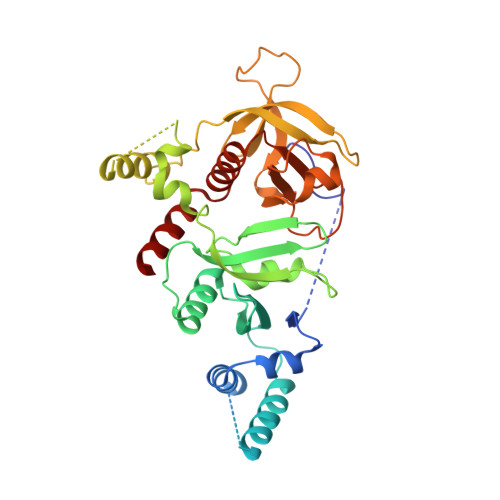
Molecular basis for autoinhibition of RIAM regulated by FAK in integrin activation.
Chang, Y.C., Su, W., Cho, E.A., Zhang, H., Huang, Q., Philips, M.R., Wu, J.(2019) Proc Natl Acad Sci U S A 116: 3524-3529
- PubMed: 30733287 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1818880116
- Primary Citation Related Structures:
6E31 - PubMed Abstract:
RAP1-interacting adapter molecule (RIAM) mediates RAP1-induced integrin activation. The RAS-association (RA) segment of the RA-PH module of RIAM interacts with GTP-bound RAP1 and phosphoinositol 4,5 bisphosphate but this interaction is inhibited by the N-terminal segment of RIAM. Here we report the structural basis for the autoinhibition of RIAM by an intramolecular interaction between the IN region (aa 27-93) and the RA-PH module. We solved the crystal structure of IN-RA-PH to a resolution of 2.4-Å. The structure reveals that the IN segment associates with the RA segment and thereby suppresses RIAM:RAP1 association. This autoinhibitory configuration of RIAM can be released by phosphorylation at Tyr45 in the IN segment. Specific inhibitors of focal adhesion kinase (FAK) blocked phosphorylation of Tyr45, inhibited stimulated translocation of RIAM to the plasma membrane, and inhibited integrin-mediated cell adhesion in a Tyr45-dependent fashion. Our results reveal an unusual regulatory mechanism in small GTPase signaling by which the effector molecule is autoinhibited for GTPase interaction, and a modality of integrin activation at the level of RIAM through a FAK-mediated feedforward mechanism that involves reversal of autoinhibition by a tyrosine kinase associated with integrin signaling.
- Molecular Therapeutics Program, Fox Chase Cancer Center, Philadelphia, PA 19111.
Organizational Affiliation: