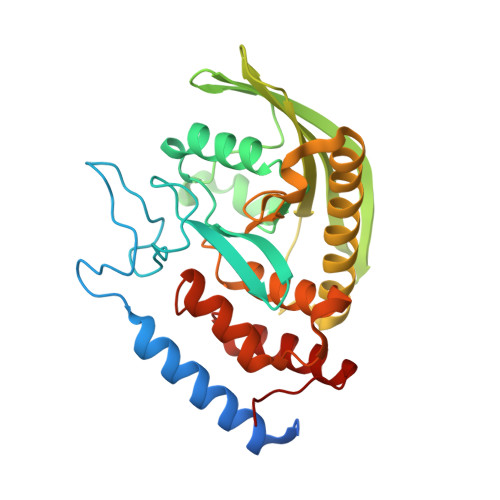
A YopH PTP1B Chimera Shows the Importance of the WPD-Loop Sequence to the Activity, Structure, and Dynamics of Protein Tyrosine Phosphatases.
Moise, G., Morales, Y., Beaumont, V., Caradonna, T., Loria, J.P., Johnson, S.J., Hengge, A.C.(2018) Biochemistry 57: 5315-5326
- PubMed: 30110154 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.8b00663
- Primary Citation Related Structures:
6DR1, 6DR7, 6DR9, 6DRB, 6DT6 - PubMed Abstract:
To study factors that affect WPD-loop motion in protein tyrosine phosphatases (PTPs), a chimera of PTP1B and YopH was created by transposing the WPD loop from PTP1B to YopH. Several subsequent mutations proved to be necessary to obtain a soluble, active enzyme. That chimera, termed chimera 3, retains productive WPD-loop motions and general acid catalysis with a pH dependency similar to that of the native enzymes. Kinetic isotope effects show the mechanism and transition state for phosphoryl transfer are unaltered. Catalysis of the chimera is slower than that of either of its parent enzymes, although its rate is comparable to those of most native PTPs. X-ray crystallography and nuclear magnetic resonance were used to probe the structure and dynamics of chimera 3. The chimera's structure was found to sample an unproductive hyper-open conformation of its WPD loop, a geometry that has not been observed in either of the parents or in other native PTPs. The reduced catalytic rate is attributed to the protein's sampling of this conformation in solution, reducing the fraction in the catalytically productive loop-closed conformation.
- Department of Chemistry and Biochemistry , Utah State University , Logan , Utah 84322-0300 , United States.
Organizational Affiliation: