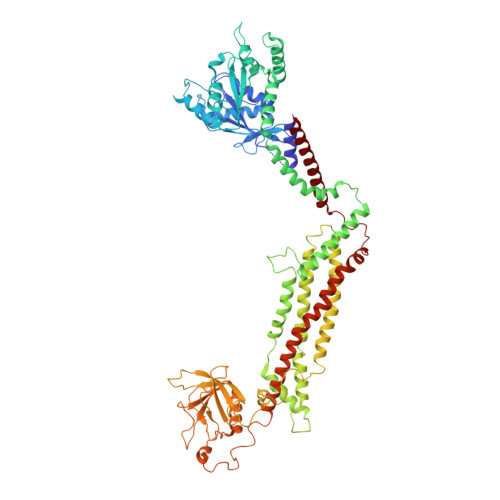
Cryo-EM of the dynamin polymer assembled on lipid membrane.
Kong, L., Sochacki, K.A., Wang, H., Fang, S., Canagarajah, B., Kehr, A.D., Rice, W.J., Strub, M.P., Taraska, J.W., Hinshaw, J.E.(2018) Nature 560: 258-262
- PubMed: 30069048 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-018-0378-6
- Primary Citation Related Structures:
6DLU, 6DLV - PubMed Abstract:
Membrane fission is a fundamental process in the regulation and remodelling of cell membranes. Dynamin, a large GTPase, mediates membrane fission by assembling around, constricting and cleaving the necks of budding vesicles 1 . Here we report a 3.75 Å resolution cryo-electron microscopy structure of the membrane-associated helical polymer of human dynamin-1 in the GMPPCP-bound state. The structure defines the helical symmetry of the dynamin polymer and the positions of its oligomeric interfaces, which were validated by cell-based endocytosis assays. Compared to the lipid-free tetramer form 2 , membrane-associated dynamin binds to the lipid bilayer with its pleckstrin homology domain (PHD) and self-assembles across the helical rungs via its guanine nucleotide-binding (GTPase) domain 3 . Notably, interaction with the membrane and helical assembly are accommodated by a severely bent bundle signalling element (BSE), which connects the GTPase domain to the rest of the protein. The BSE conformation is asymmetric across the inter-rung GTPase interface, and is unique compared to all known nucleotide-bound states of dynamin. The structure suggests that the BSE bends as a result of forces generated from the GTPase dimer interaction that are transferred across the stalk to the PHD and lipid membrane. Mutations that disrupted the BSE kink impaired endocytosis. We also report a 10.1 Å resolution cryo-electron microscopy map of a super-constricted dynamin polymer showing localized conformational changes at the BSE and GTPase domains, induced by GTP hydrolysis, that drive membrane constriction. Together, our results provide a structural basis for the mechanism of action of dynamin on the lipid membrane.
- Laboratory of Cell and Molecular Biology, National Institute of Diabetes and Digestive and Kidney Diseases, NIH, Bethesda, MD, USA.
Organizational Affiliation: