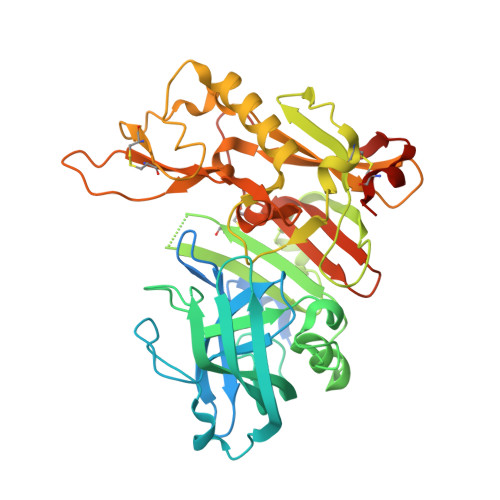
Design, synthesis, X-ray studies, and biological evaluation of novel BACE1 inhibitors with bicyclic isoxazoline carboxamides as the P3 ligand.
Ghosh, A.K., Ghosh, K., Brindisi, M., Lendy, E.K., Yen, Y.C., Kumaragurubaran, N., Huang, X., Tang, J., Mesecar, A.D.(2018) Bioorg Med Chem Lett 28: 2605-2610
- PubMed: 29970308 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bmcl.2018.06.045
- Primary Citation Related Structures:
6DHC - PubMed Abstract:
We describe the design, synthesis, X-ray studies, and biological evaluation of novel BACE1 inhibitors containing bicyclic isoxazoline carboxamides as the P3 ligand in combination with methyl cysteine, methylsulfonylalanine and Boc-amino alanine as P2 ligands. Inhibitor 3a displayed a BACE1 K i value of 10.9 nM and EC 50 of 343 nM. The X-ray structure of 3a bound to the active site of BACE1 was determined at 2.85 Å resolution. The structure revealed that the major molecular interactions between BACE1 and the bicyclic tetrahydrofuranyl isoxazoline heterocycle are van der Waals in nature.
- Department of Chemistry, Purdue University, West Lafayette, IN 47907, United States; Department of Medicinal Chemistry, Purdue University, West Lafayette, IN 47907, United States. Electronic address: akghosh@purdue.edu.
Organizational Affiliation: