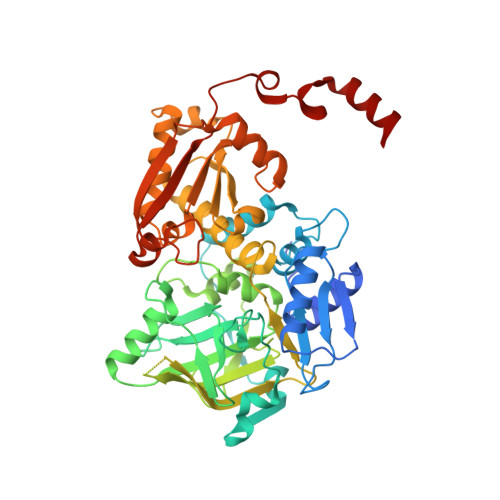
Biochemical and Structural Characterization of TtnD, a Prenylated FMN-Dependent Decarboxylase from the Tautomycetin Biosynthetic Pathway.
Annaval, T., Han, L., Rudolf, J.D., Xie, G., Yang, D., Chang, C.Y., Ma, M., Crnovcic, I., Miller, M.D., Soman, J., Xu, W., Phillips Jr., G.N., Shen, B.(2018) ACS Chem Biol 13: 2728-2738
- PubMed: 30152678
- DOI: https://doi.org/10.1021/acschembio.8b00673
- Primary Citation Related Structures:
6DA6, 6DA7, 6DA9 - PubMed Abstract:
Tautomycetin (TTN) is a polyketide natural product featuring a terminal alkene. Functional characterization of the genes within the ttn gene cluster from Streptomyces griseochromogenes established the biosynthesis of the TTN polyketide backbone, its dialkylmaleic anhydride moiety, the coupling of the two moieties to form the nascent intermediate TTN F-1, and the tailoring steps converting TTN F-1 to TTN. Here, we report biochemical and structural characterization of TtnD, a prenylated FMN (prFMN)-dependent decarboxylase belonging to the UbiD family that catalyzes the penultimate step of TTN biosynthesis. TtnD catalyzes decarboxylation of TTN D-1 to TTN I-1, utilizing prFMN as a cofactor generated by the TtnC flavin prenyltransferase; both TtnD and TtnC are encoded within the ttn biosynthetic gene cluster. TtnD exhibits substrate promiscuity but accepts only TTN D-1 congeners that feature an α,β-unsaturated acid, supporting the [3+2] cycloaddition mechanism during catalysis that requires the double bond of an α,β-unsaturated acid substrate. TtnD shares a similar overall structure with other members of the UbiD family but forms a homotetramer in solution. Each protomer is composed of three domains with the active site located between the middle and C-terminal domains; R169-E272-E277, constituting the catalytic triad, and E228, involved in Mn(II)-mediated binding of prFMN, were confirmed by site-directed mutagenesis. TtnD represents the first example of a prFMN-dependent decarboxylase involved in polyketide biosynthesis, expanding the substrate scope of the UbiD family of decarboxylases beyond simple aromatic and cinnamic acids. TtnD and its homologues are widespread in nature and could be exploited as biocatalysts for organic synthesis.
- Department of Chemistry , The Scripps Research Institute , 130 Scripps Way , Jupiter , Florida 33458 , United States.
Organizational Affiliation: