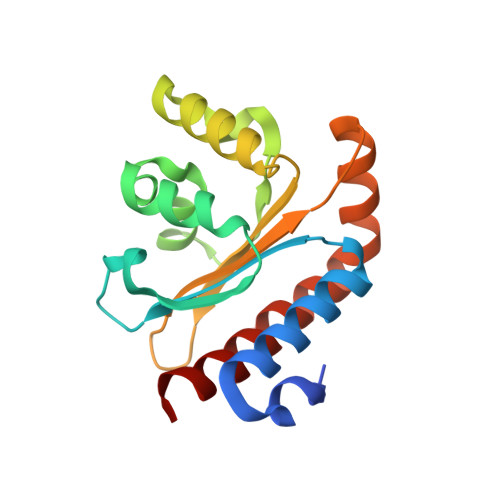
Structural and Biochemical Studies of Non-native Agonists of the LasR Quorum-Sensing Receptor Reveal an L3 Loop "Out" Conformation for LasR.
O'Reilly, M.C., Dong, S.H., Rossi, F.M., Karlen, K.M., Kumar, R.S., Nair, S.K., Blackwell, H.E.(2018) Cell Chem Biol 25: 1128-1139.e3
- PubMed: 30033130
- DOI: https://doi.org/10.1016/j.chembiol.2018.06.007
- Primary Citation of Related Structures:
6D6A, 6D6B, 6D6C, 6D6D, 6D6L, 6D6M, 6D6N, 6D6O, 6D6P - PubMed Abstract:
Chemical strategies to block quorum sensing (QS) could provide a route to attenuate virulence in bacterial pathogens. Considerable research has focused on this approach in Pseudomonas aeruginosa, which uses the LuxR-type receptor LasR to regulate much of its QS network. Non-native ligands that antagonize LasR have been developed, yet we have little understanding of the mode by which these compounds interact with LasR and alter its function, as the receptor is unstable in their presence. Herein, we report an approach to circumvent this challenge through the study of a series of synthetic LasR agonists with varying levels of potency. Structural investigations of these ligands with the LasR ligand-binding domain reveal that certain agonists can enforce a conformation that deviates from that observed for other, often more potent agonists. These results, when combined with cell-based and biophysical analyses, suggest a functional model for LasR that could guide future ligand design.
- Department of Chemistry, University of Wisconsin-Madison, 1101 University Avenue, Madison, WI 53706, USA.
Organizational Affiliation: