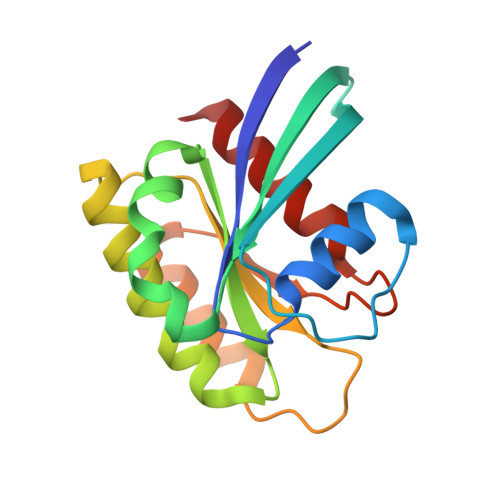
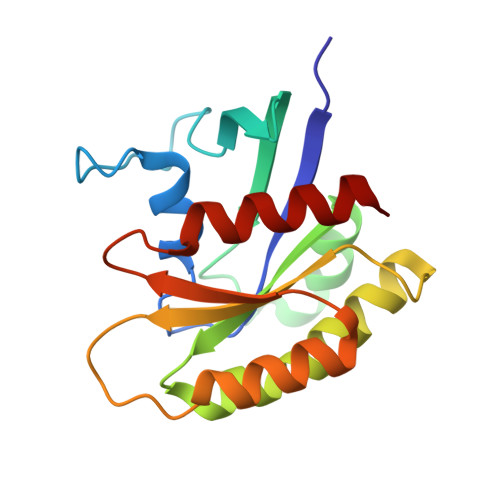
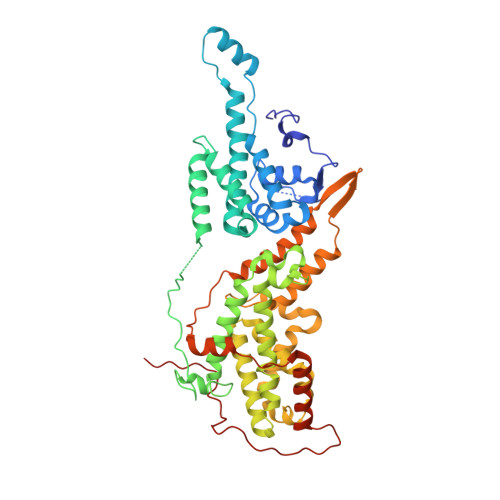
Discovery and Structure-Based Optimization of Benzimidazole-Derived Activators of SOS1-Mediated Nucleotide Exchange on RAS.
Hodges, T.R., Abbott, J.R., Little, A.J., Sarkar, D., Salovich, J.M., Howes, J.E., Akan, D.T., Sai, J., Arnold, A.L., Browning, C., Burns, M.C., Sobolik, T., Sun, Q., Beesetty, Y., Coker, J.A., Scharn, D., Stadtmueller, H., Rossanese, O.W., Phan, J., Waterson, A.G., McConnell, D.B., Fesik, S.W.(2018) J Med Chem 61: 8875-8894
- PubMed: 30205005 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.jmedchem.8b01108
- Primary Citation Related Structures:
6D55, 6D56, 6D59, 6D5E, 6D5G, 6D5H, 6D5J, 6D5L, 6D5M, 6D5V, 6D5W - PubMed Abstract:
Son of sevenless homologue 1 (SOS1) is a guanine nucleotide exchange factor that catalyzes the exchange of GDP for GTP on RAS. In its active form, GTP-bound RAS is responsible for numerous critical cellular processes. Aberrant RAS activity is involved in ∼30% of all human cancers; hence, SOS1 is an attractive therapeutic target for its role in modulating RAS activation. Here, we describe a new series of benzimidazole-derived SOS1 agonists. Using structure-guided design, we discovered small molecules that increase nucleotide exchange on RAS in vitro at submicromolar concentrations, bind to SOS1 with low double-digit nanomolar affinity, rapidly enhance cellular RAS-GTP levels, and invoke biphasic signaling changes in phosphorylation of ERK 1/2. These compounds represent the most potent series of SOS1 agonists reported to date.
- Boehringer Ingelheim RCV GmbH & Co. KG , Doktor-Boehringer-Gasse 5-11 , 1120 Vienna , Austria.
Organizational Affiliation: