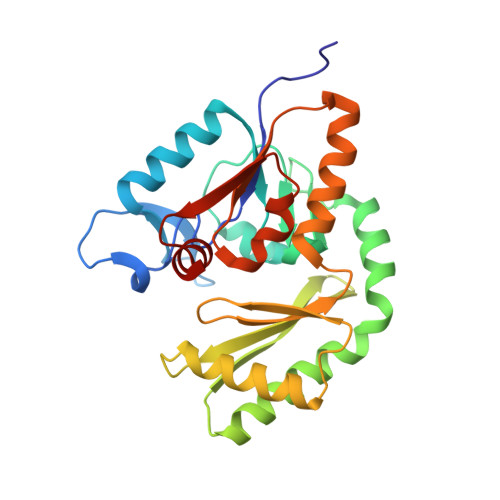
Trehalose 6-phosphate phosphatases of Pseudomonas aeruginosa.
Cross, M., Biberacher, S., Park, S.Y., Rajan, S., Korhonen, P., Gasser, R.B., Kim, J.S., Coster, M.J., Hofmann, A.(2018) FASEB J 32: 5470-5482
- PubMed: 29688811
- DOI: https://doi.org/10.1096/fj.201800500R
- Primary Citation Related Structures:
6CJ0, 6D3V, 6D3W - PubMed Abstract:
The opportunistic bacterium Pseudomonas aeruginosa has been recognized as an important pathogen of clinical relevance and is a leading cause of hospital-acquired infections. The presence of a glycolytic enzyme in Pseudomonas, which is known to be inhibited by trehalose 6-phosphate (T6P) in other organisms, suggests that these bacteria may be vulnerable to the detrimental effects of intracellular T6P accumulation. In the present study, we explored the structural and functional properties of trehalose 6-phosphate phosphatase (TPP) in P. aeruginosa in support of future target-based drug discovery. A survey of genomes revealed the existence of 2 TPP genes with either chromosomal or extrachromosomal location. Both TPPs were produced as recombinant proteins, and characterization of their enzymatic properties confirmed specific, magnesium-dependent catalytic hydrolysis of T6P. The 3-dimensional crystal structure of the chromosomal TPP revealed a protein dimer arising through β-sheet expansion of the individual monomers, which possess the overall fold of halo-acid dehydrogenases.-Cross, M., Biberacher, S., Park, S.-Y., Rajan, S., Korhonen, P., Gasser, R. B., Kim, J.-S., Coster, M. J., Hofmann, A. Trehalose 6-phosphate phosphatases of Pseudomonas aeruginosa.
- Griffith Institute for Drug Discovery, Griffith University, Nathan, Queensland, Australia.
Organizational Affiliation: