Structural and immunologic correlates of chemically stabilized HIV-1 envelope glycoproteins.
Schiffner, T., Pallesen, J., Russell, R.A., Dodd, J., de Val, N., LaBranche, C.C., Montefiori, D., Tomaras, G.D., Shen, X., Harris, S.L., Moghaddam, A.E., Kalyuzhniy, O., Sanders, R.W., McCoy, L.E., Moore, J.P., Ward, A.B., Sattentau, Q.J.(2018) PLoS Pathog 14: e1006986-e1006986
- PubMed: 29746590 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1006986
- Primary Citation Related Structures:
6CRQ - PubMed Abstract:
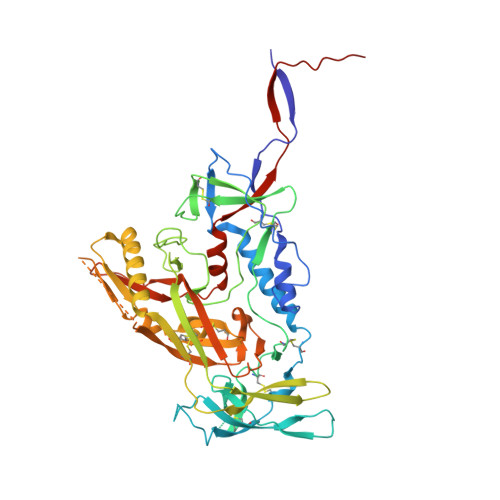
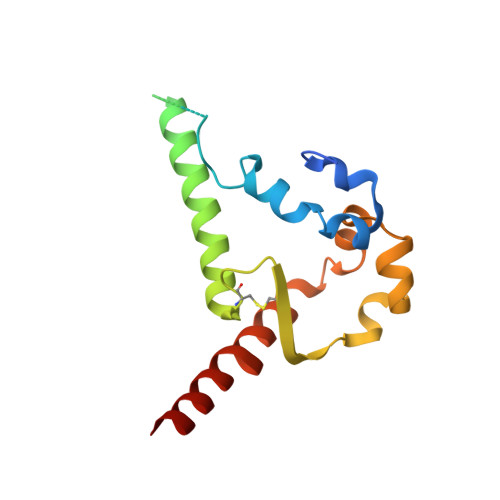
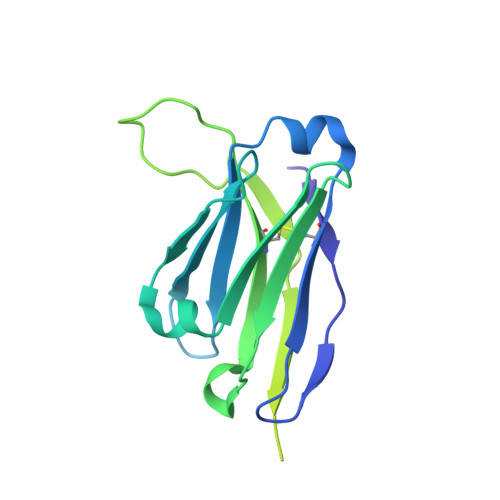
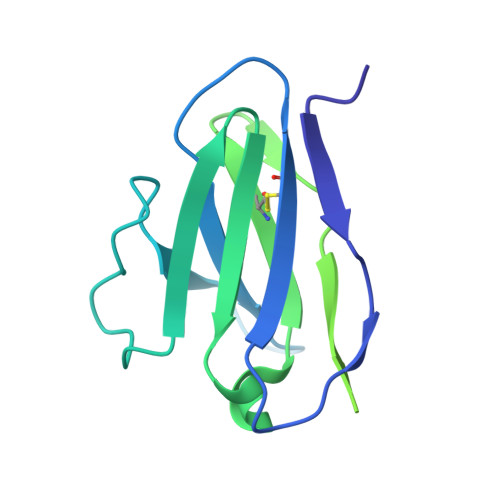
Inducing broad spectrum neutralizing antibodies against challenging pathogens such as HIV-1 is a major vaccine design goal, but may be hindered by conformational instability within viral envelope glycoproteins (Env). Chemical cross-linking is widely used for vaccine antigen stabilization, but how this process affects structure, antigenicity and immunogenicity is poorly understood and its use remains entirely empirical. We have solved the first cryo-EM structure of a cross-linked vaccine antigen. The 4.2 Å structure of HIV-1 BG505 SOSIP soluble recombinant Env in complex with a CD4 binding site-specific broadly neutralizing antibody (bNAb) Fab fragment reveals how cross-linking affects key properties of the trimer. We observed density corresponding to highly specific glutaraldehyde (GLA) cross-links between gp120 monomers at the trimer apex and between gp120 and gp41 at the trimer interface that had strikingly little impact on overall trimer conformation, but critically enhanced trimer stability and improved Env antigenicity. Cross-links were also observed within gp120 at sites associated with the N241/N289 glycan hole that locally modified trimer antigenicity. In immunogenicity studies, the neutralizing antibody response to cross-linked trimers showed modest but significantly greater breadth against a global panel of difficult-to-neutralize Tier-2 heterologous viruses. Moreover, the specificity of autologous Tier-2 neutralization was modified away from the N241/N289 glycan hole, implying a novel specificity. Finally, we have investigated for the first time T helper cell responses to next-generation soluble trimers, and report on vaccine-relevant immunodominant responses to epitopes within BG505 that are modified by cross-linking. Elucidation of the structural correlates of a cross-linked viral glycoprotein will allow more rational use of this methodology for vaccine design, and reveals a strategy with promise for eliciting neutralizing antibodies needed for an effective HIV-1 vaccine.
- The Sir William Dunn School of Pathology, The University of Oxford, Oxford, United Kingdom.
Organizational Affiliation: